Phytoplasmas, previously termed mycoplasma-like organisms, are small, cell wall-less, phloem-limited prokaryotes that have been reported to be associated with diseases in several hundred to more than 1 000 plant species worldwide (Lee, Davis, & Gundersen-Rindal, 2000; Hogenhout, et al., 2008). They were first discovered, through electron microscopy studies, by a group of Japanese scientists in 1967 (Doi, Teranaka, Yora, & Asuyama, 1967). Based on phylogenetic analyses using 16S rRNA and/or ribosomal protein gene sequences, phytoplasmas have been placed in the class Mollicutes (Gundersen, Lee, Rehner, Davis, & Kingsbury, 1994). Phytoplasma infection interferes with plant development and induces morphological and physiological changes, including: witches’-broom, phyllody, virescence, bolting, reddening of leaves and stems, generalized yellowing, stunting and decline (Hogenhout et al., 2008). These mollicutes are responsible for devastating damage to many economically important crops, fruit trees, woody trees and ornamental plants worldwide. Some of these diseases can cause death of the plant host resulting in serious economic impact and may affect local biodiversity (Maejima, Kenro Oshima, & Namda, 2014; Marcone, 2014).
In nature, phytoplasmas live and reproduce in the phloem tissue of plants, as well as in the salivary glands and other tissues of some phloem feeding Hemiptera of the families Cicadellidae, Cixiidae, Delphacidae, Derbidae, Flatidae and Psyllidae. Phytoplasmal diseases are transmitted from plant to plant by one of such insect vectors, as well as by grafting and Cuscuta spp. A suitable combination of various plant hosts and insect vector-mediated transmission are responsible for the horizontal transmission of phytoplasmas between plants (Weintraub & Beanland, 2006). Some studies suggest the possibility of transovarial transmission of phytoplasmas (Alma et al., 1997; Hanboonsong, Choosai, Panyim, & Damak, 2002; Mittelberger et al., 2017).
Phytoplasmas are difficult to cultivate in vitro, and are therefore poorly characterized bacteria. Molecular methods using the highly conserved 16S rRNA gene as well as other conserved biomarkers, including ribosomal protein, tuf, secA, and secY genes, have been used for the detection, differentiation, and classification of phytoplasmas (Lee, Gundersen-Rindal, Davis, & Batoszyk, 1998b; Seemüller, Marcone, Lauer, Ragozzino, & Göschl, 1998; Marcone, Lee, Davis, Ragozzino, & Seemüller, 2000; Hodgetts, Boonham, Mumford, Harrison, & Dickinson, 2008). Based on molecular classification and the guidelines established by the IRPCM Phytoplasma/Spiroplasma Working Team-Phytoplasma Taxonomy Group (IRPCM, 2004), phytoplasmas have been classified into 43 ‘Candidatus Phytoplasma’ species (Arneodo, et al., 2007; Fernández, Galdeano, Kornowski, Arneado, & Conci, 2016; Zhao & Davis, 2016; Miyazaki et al., 2017; Naderali et al., 2017).
Phytoplasma infection with one of four 16Sr groups (16SrI, 16SrIII, 16SrIX, and 16Sr XV) has been reported in various plant species in Costa Rica (Gámez & León, 1985; Kenyon, Harrison, & Richardson, 1999; Villalobos, Moreira, Bottner, Lee, & Rivera, 2002; Villalobos et al., 2011; Pardo, Truke, Cardozo, Varela, & Alvarez, 2014). Although natural phytoplasma infection in Catharanthus roseus G. Don (Apocynaceae) has been reported elsewhere (Pérez-López, Olivier, Luna-Rodríguez, Adame-García, & Dumonceaux, 2016b), to our knowledge, it has not been reported in Costa Rica. This plant species, a perennial herbaceous plant native to Madagascar, is commonly known as Madagascar periwinkle, vinca, or “mariposa” (butterfly) in Costa Rica. It is widely cultivated as a popular ornamental plant and can be found in gardens and homes across the warmer parts of tropical and subtropical countries. Periwinkle plants are used as an experimental host for the maintenance of phytoplasma strains, as well as to study phytoplasma-host interactions (Nejat et al., 2013; 2015).
We consistently found Madagascar periwinkle plants exhibiting symptoms reminiscent of phytoplasma infection (Hogenhout et al., 2008) in gardens, parks, living-fences and along sidewalks throughout Costa Rica. In this study, we examined a sample of 73 symptomatic C. roseus plants collected from 2012 to 2016. The phytoplasmas were detected and identified by nested PCR, sequencing, in silico RFLP and phylogenetic analyses. This work reports natural infection of Madagascar periwinkle plants with different phytoplasma 16Sr subgroups in Costa Rica.
Materials and methods
Sampling: A total of 73 C. roseus plants exhibiting different symptoms related to phytoplasma infection (witches’-broom, phyllody, virescense, leaf yellowing, dwarfing, etc.) were collected from home gardens, sidewalks, and parks at different locations in the seven provinces of Costa Rica [Alajuela province: Alajuela, Cartago province: Dulce Nombre, Paraíso, Turrialba; Guanacaste province: Hojancha, Filadelfia, Cañas, Sámara, Santa Cruz; Heredia province: Santo Domingo; Limón province: Cahuita; Puntarenas province: Chomes, Esparza, Paso Canoas, Potrero Grande, Quebrada Grande, Tárcoles; and San José province: Coronado, Moravia, Pérez Zeledón, San Pedro, Sabanilla]. Two Gliricidia sepium trees showing little leaf disease (GLLD) were collected in La Guácima (Alajuela province) to compare sequences with those from C. roseus samples harboring group 16SrIX phytoplasmas.
DNA extraction and amplification: Leaf midribs and petiole (100 mg) from each sample of C. roseus and G. sepium were extracted using the DNeasy Plant Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instruction. Additionally, DNA from healthy periwinkle plants grown in an insect-proof greenhouse was extracted as negative control. DNA extracted from Sechium edule infected with 16SrI-B phytoplasma (Villalobos et al., 2002) was used as a positive control.
Detection of phytoplasmas was carried out by amplification of 16S rDNA in a nested polymerase chain reaction (PCR) assay, using universal primer pair P1/P7 (Deng & Hiruki, 1991; Smart, et al., 1996) in the first reaction followed by R16F2n/R16R2 (Gundersen, Lee, Rehner, Davis, & Kingsbury, 1994; Lee, Gundersen, Hammond, & Davis, 1994) in the second reaction. Amplifications were performed with a PCR Gradient Palm Cycler (Corbett Research Model CG1-96, Australia) in 27 μl reactions containing 200 μM of each of the four dNTPs, 0.4 μM of each primer, 1.5 mM MgCl2, 0.625 units of DreamTaq DNA polymerase (Thermo Fisher Scientific Inc. USA), and 2 μl of DNA extracted. Diluted PCR product (1:20) with UltraPure DNase/RNase free distilled water (Thermo Fisher Scientific Inc.) from the first amplification was used as template in the nested PCR. The PCR thermocycler profile for both amplification steps was denaturation at 94 °C for 1 min (2 min for the first cycle), annealing at 55 °C for 1 min, and extension at 72 °C for 2 min (10 min for the last cycle). The amplified products (5 μl of each PCR reaction) were evaluated by electrophoresis through 1 % agarose gels, stained with GelRedTM (Biotium, California, USA) and visualized with a UV transilluminator.
Sequencing and phylogenetic analyses: All samples (n = 52) that yielded amplicons of about 1.2 Kb in the previous nested PCR were used to prepare new reactions using only the internal primer pairs of the nested PCR protocol (Gundersen & Lee, 1996a) and were directly sequenced in both directions by Macrogen Inc. (Korea). A contig sequence using the forward and reverse sequences was obtained for each sample using BioEdit software v. 7.2.5 (Hall, 1999). Confirmation of phytoplasmas infection and preliminary identification of the phytoplasma group for each sample was done using BLAST algorithm (http://blast.ncbi.nlm.nih.gov/Blast.cgi).
A total of five phytoplasma groups were identified within those 52 positive samples (Digital Appendix 1). Four representative samples for group 16SrI (the most abundant) and one each from the other four phytoplasma groups (16SrIII, 16SrIX, 16SrXIII and 16SrXV) were selected for further analyses. Additionally, two samples of G. sepium were included as control for phytoplasma group 16SrIX. A semi-nested PCR assay was performed using primer pair P1/16S-SR in the first reaction followed by P1A/16S-SR (Lee, et al., 2004) in the second reaction to amplify the near full length 16S rRNA gene (about 1.5 Kb). The nested PCR products (a total of eight) were purified using PCR Kleen Spin Columns (Bio-Rad, Hercules, CA) and cloned into Escherichia coli (TOP10) by using the TOPO TA Cloning Kit (Invitrogen, Carlsbad, CA) according to the manufacturers’ instructions. Two clones per sample were sequenced with an automated DNA sequencer (Macrogen Inc., Rockville, MD USA). Sequencing reads were assembled using the SeqMan program from the DNAStar LaserGene software package (DNAStar, Madison, WI) and a final sequence was obtained for each clone. The iPhyClassifier tool (Zhao, et al., 2009) was then used to generate in silico RFLP profiles and determine phytoplasma group classifications. A BLAST search (BLASTn) was utilized to find similar sequences deposited in GenBank. Sequences from clones obtained from C. roseus have been deposited to GenBank under accession numbers MH428957 to MH428964. Additionally, sequences of Costa Rican GLLD are available as MH428965 and MH428966.
The sequences of subgroup representatives obtained in this study as well as 30 phytoplasma strains available from GenBank, and Acholeplasma palmae J233 (as outgroup) were aligned using ClustalW (Thompson, Higgins, & Gibson, 1994) in MEGA v. 7.0 (Kumar, Stecher, & Tamura, 2016). The final alignment included a total of 1 401 positions. The phylogenetic analysis was inferred with MEGA7 using the Maximum Likelihood method based on the General Time Reversible model and the rate of variation per site was determined by a gamma distribution with a proportion of invariable sites (G+I). The estimation of stability and support for the inferred clades were performed using bootstrap analyses of 2 500 replicates.
Results
A total of 73 C. roseus samples were collected in different regions of the country and phytoplasma infection was detected in 71.2 % (52/73) by nested PCR. Ten different symptom combinations (Table 1) were observed in the phytoplasma-positive C. roseus plants (Figure 1A, Figure 1B, Figure 1C, Figure 1D, Figure 1E and Figure 1F). The most common symptom was virescence (19 plants) or virescence combined with other symptoms (11 plants), representing in total 57.7 % of the positive plants. The second most abundant symptom was yellowing, observed on 19 plants (36.5 %). We were not able to find a pattern/association between type of symptoms and other variables, including phytoplasma group or geographic altitude where the plant was collected. Nonetheless, perhaps due to a greater number of samples, the 16SrI group showed variation regarding symptom expression in the plant host C. roseus.
Table 1: Number of samples in each symptom category associated with each of five phytoplasma 16Sr groups identified in Catharanthus roseus collected in Costa Rica from 2012-2016. Cuadro 1: Número de muestras según síntoma observado en Catharanthus roseus infectados con fitoplasmas, recolectadas en Costa Rica, 2012-2016
| Symptoms / Síntomas | Phytoplasma group/ Grupo de fitoplasma | Total | ||||
| 16SrI | 16SrIII | 16SrIX | 16SrXIII | 16SrXV | ||
| Bolting | 1 | - | - | - | - | 1 |
| Phyllody + little leaf + proliferation | 2 | - | - | - | - | 2 |
| Virescence | 14 | 2 | 1 | 1 | 1 | 19 |
| Virescence + big bud | - | - | - | - | 1 | 1 |
| Virescence + phyllody | 1 | - | - | - | 1 | 2 |
| Virescence + proliferation | 1 | - | - | - | - | 1 |
| Virescence + proliferation + little leaf | 2 | - | - | 1 | - | 3 |
| Virescence + little leaf + stunting + short internodes | - | 3 | - | - | - | 3 |
| Virescence + yellowing | 1 | - | - | - | - | 1 |
| Yellowing | 8 | 9 | 2 | - | - | 19 |
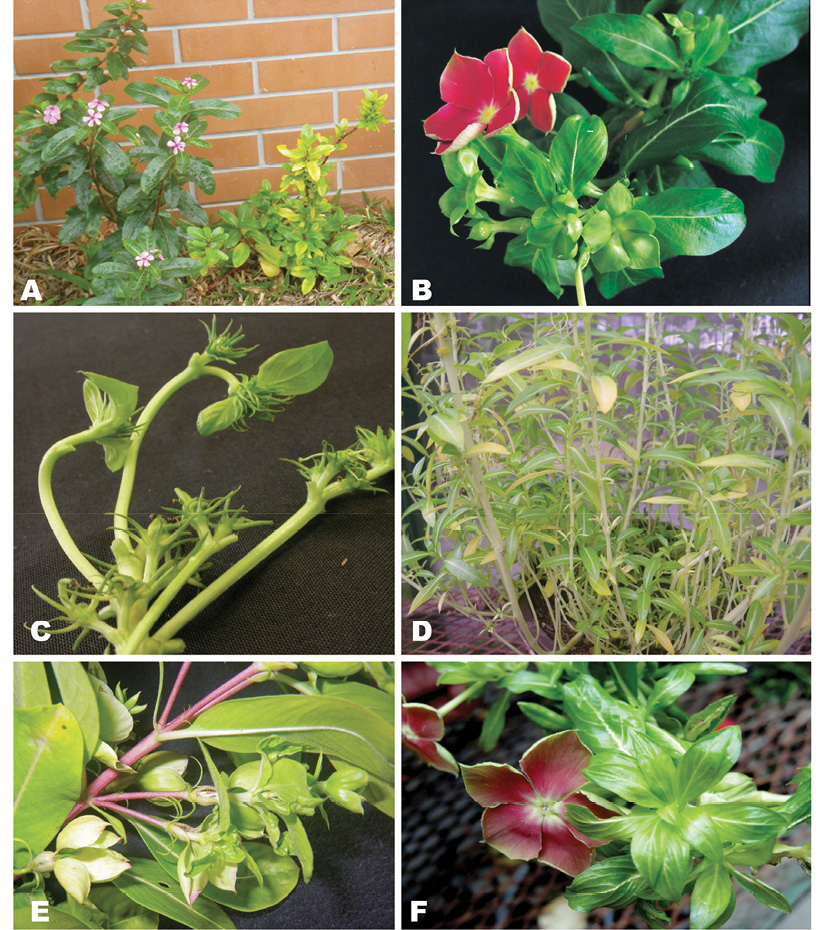

Figure 1 Symptoms exhibited by periwinkle plants infected with phytoplasmas including A) yellowing, dwarfing, little leaf and axilar proliferation, B) virescence, C) floral abortion and malformation, D) stalk elongation (bolting), E) big bud, F) phyllody.
According to BLAST search results, phytoplasmas belonging to groups: 16SrI, 16SrIII, 16SrIX, 16SrXIII and 16SrXV were detected in 30, 14, 3, 2 and 3 symptomatic C. roseus samples (Table 2), respectively. No mixed infections were found, despite the presence of two or three different phytoplasmas in some locations in the same province (Table 2). The altitudinal distribution of phytoplasmas suggested that 16SrI group has a wide distribution, while groups 16SrIII and 16SrXV showed presence above 1 200 masl or below 500 masl, respectively (Figure 2).
Table 2: Number of infected Catharanthus roseus samples associated with each of five phytoplasma 16Sr groups identified and province of collection in Costa Rica. Cuadro 2: Número de muestras de Catharanthus roseus según grupo de fitoplasma detectado por provincia de recolección en Costa Rica
| Province / Provincia | Phytoplasma group / Grupo de fitoplasma | Total | ||||
| 16SrI | 16SrIII | 16SrIX | 16SrXIII | 16SrXV | ||
| Cartago | 7 | 1 | - | 1 | - | 9 |
| Guanacaste | 6 | - | - | 1 | 3 | 10 |
| Heredia | 1 | 1 | - | - | - | 2 |
| Limón | 1 | - | - | - | - | 1 |
| Puntarenas | 4 | - | 1 | - | - | 5 |
| San José | 11 | 12 | 2 | - | - | 25 |
| Total | 30 | 14 | 3 | 2 | 3 | 52 |
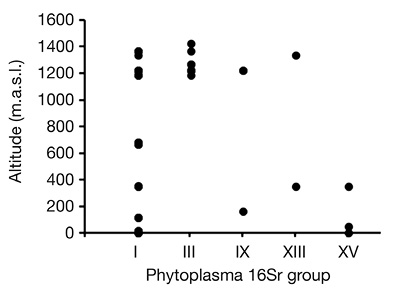
Figure 2 Altitudinal distribution (meters above sea level, m.a.s.l.) of Catharantus roseus samples per phytoplasma group collected in this study in Costa Rica.
The majority of nucleotide sequences obtained from positive plants (30 of 52 sequences) indicated infection with ‘Ca. Phytoplasma asteris’, with similarities between 98-99 % with subgroup 16SrI-B representatives. Plants infected with this subgroup were found in six out of seven provinces in the country (Table 2). Only one infected C. roseus (CR02) expressing bolting symptoms (GenBank accession no. MH428961), detected in Turrialba city (Cartago province), showed 97 % similarity to subgroup 16SrI-P (GenBank accession no. AF503568).
A summary of data obtained using the iPhyClassifier tool to analyze cloned phytoplasma 16S rRNA gene sequences obtained in this study is presented in Table 3. The virtual RFLPs patterns of three sequences representative of subgroup 16SrI-B (GenBank accession nos. MH428957, MH428960 and MH428964) were identical to the reference strain of this subgroup. However, the sequence of a sample from Turrialba identified as subgroup 16SrI-P (GenBank accession no. MH428961) showed identical virtual RFLP patterns to this subgroup with 16 out of 17 enzymes used, but the pattern resolved using HhaI was like other strains in the 16SrI group although dissimilar to subgroup 16SrI-P (Figure 3A), suggesting a possible new subgroup within the ‘Ca. Phytoplasma asteris’ group in Costa Rica.
Table 3: Summary of data obtained using iPhyClassifier tool to analyze cloned phytoplasmas 16S rRNA sequences obtained in this study. Cuadro 3: Resumen del análisis de datos para las secuencias de fitoplasmas obtenidos en este estudio usando la herramienta iPhyClassifier
| Sample ID | Gen Bank Accession number | ‘CandidatusPhytoplasma sp.', assignment using reference strains GenBank accession numbers indicated in parentheses | Sequence similarity to the assigned ‘Ca. Phytoplasma sp.' reference strain (%) | Similarity coefficient to the reference pattern of phytoplasma 16Sr group/subgroup (GenBank accession number) | Relation to 16Sr ubgroup / group classified | Restriction enzyme delineating |
| CR78 | MH428957 | ‘Ca. Phytoplasma asteris' (M30790) | 99.5 | 1.00, 16Sr I-B (AP006628) | 16SrI-B member | Virtual RFLP patterns identical to 16SrI-B |
| CR41 | MH428960 | ‘Ca. Phytoplasma asteris' (M30790) | 99.5 | 1.00, 16Sr I-B (AP006628) | 16SrI-B member | Virtual RFLP patterns identical to 16SrI-B |
| CR47 | MH428964 | ‘Ca. Phytoplasma asteris' (M30790) | 99.6 | 1.00, 16Sr I-B (AP006628) | 16SrI-B member | Virtual RFLP patterns identical to 16SrI-B |
| CR02 | MH428961 | ‘Ca. Phytoplasma asteris' (M30790) | 99.6 | 0.97, 16SrI-P (AF503568) | tentative 16SrI new subgroup | HhaI, pattern correspond to 16SrI-B instead of 16SrI-P |
| CR33 | MH428959 | 'Ca. Phytoplasma pruni' rrnA’ (JQ044393) | 99.1 | 0.98, 16SrIII-F (AF510724) | 16SrIII-F variant | BstUI, pattern correspond to 16SrIII-D instead of 16SrIII-F |
| CR06 | MH428962 | ‘Ca. Phytoplasma phoenicium' (AF515636) | 98.4 | 0.97, 16SrIX-A (AF248957) | tentative 16SrIX new subgroup | RsaI, new pattern |
| GLLDCR1 | MH428965 | ‘Ca. Phytoplasma phoenicium' (AF515636) | 98.3 | 1.00, 16SrIX-F (AF361017) | 16SrIX-F member | Virtual RFLP patterns identical to 16SrIX-F |
| GLLDCR2 | MH428966 | ‘Ca. Phytoplasma phoenicium' (AF515636) | 98.3 | 1.00, 16SrIX-F (AF361017) | 16SrIX-F member | Virtual RFLP patterns identical to 16SrIX-F |
| CR25 | MH428958 | ‘Ca. Phytoplasma hispanicum' (AF248960) | 99.5 | 0.98, 16SrXIII-H (JX626329) | 16SrXIII-H variant | MseI, pattern correspond to 16SrXIII-A instead of 16SrXIII-H |
| CR14 | MH428963 | ‘Ca. Phytoplasma brasiliense' (AF147708) | 99.2 | 0.96; 16SrXV-A (AF147708) | tentative 16SrXV new subgroup | HaeIII, new pattern |
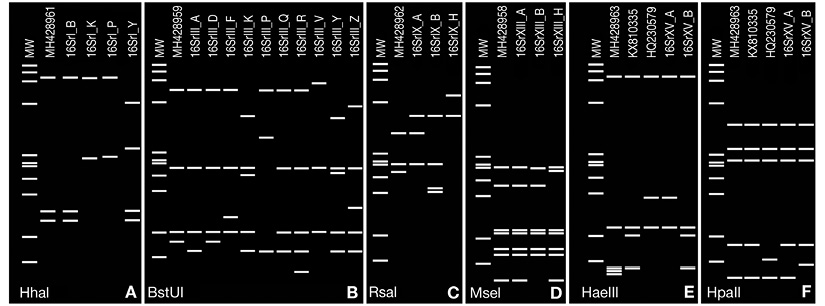
Figure 3 Computer-simulated virtual RFLP patterns generated from in silico digestions of phytoplasma 16S rRNA genes from strains detected in this study belonging to groups: (A) 16SrI using HhaI (MH428961: 16SrI-P*); (B) 16SrIII using BstUI (MH428959: 16SrIII-F*); (C) 16SrIX using RsaI (MH428962:16SrIX-A∞); (D) 16SrXIII using MseI (MH428958: 16SrXIII-H*); (E, F) 16SrXV using HaeIII and HpaII (MH428963: 16SrXV-A∞), obtained using iPhyClassifier tool. GenBank accession numbers are shown in parenthesis after each subgroup. Phytoplasma subgroup variant are indicated by an asterisk and tentative new subgroup by ∞ symbol after subgroup number. GenBank acc.nos. KX810335 and HQ230579 in E and F correspond to other variants to subgroups 16SrXV-A and 16SrXV-B reported previously. Lanes MW, HaeIII digest of ΦX174 RFI DNA; fragment sizes (bp) from top to bottom: 1353, 1078, 872, 603, 310, 281, 271, 234, 194, 118, 72. Figure edited with Paint.net software.
The second group in importance, by number of detections, was group 16SrIII (X-disease group or ‘Ca. Phytoplasma pruni’). Eleven samples collected from gardens in three different provinces (Heredia, Cartago and San José) were infected with group 16SrIII phytoplasma strains. The search for similar sequences in public databases showed 99 % identity to Mexican Xalapa periwinkle virescence phytoplasma (GenBank accession no. KY778009). The virtual RFLP pattern derived from one of our samples (GenBank accession no. MH428959) was most similar to the reference pattern of the 16SrIII-F subgroup (GenBank accession no. AF510724), with a similarity coefficient of 0.98. However, the MH428959 phytoplasma is a variant of subgroup 16SrIII-F, because the virtual RFLP pattern resolved by BstUI corresponded to subgroup 16SrIII-D instead of 16SrIII-F (Figure 3B).
The presence of three other groups: 16SrIX (‘Ca. Phytoplasma phoenicium’), 16Sr-XIII (‘Ca. Phytoplasma hispanicum’) and 16Sr-XV (‘Ca. Phytoplasma brasiliense’) was detected in a limited number of samples. Only eight plants in total from three provinces (Cartago, Guanacaste, and San José) carried one of these groups. Three C. roseus samples infected with 16SrIX group phytoplasmas were detected. A sample collected in Potrero Grande (Puntarenas), GenBank accession no. MH428962, showed 99 % similarity to pigeon pea witches’-broom (GenBank accession no. KJ817866), Crotalaria juncea witches’-broom (GenBank accession no. KF941131), and other phytoplasmal strains belonging to that group. Results from the iPhyclassifier tool for this sample classified it (similarity coefficient = 0.97) as belonging to subgroup 16SrIX-A (PPWB, GenBank accession: AF248957). However, the virtual RFLPs obtained for this sample displayed a unique RFLP pattern of three bands with enzyme RsaI (Figure 3C). When this sample was compared to sequences obtained in Costa Rica during this research from GLLD, GenBank accession nos. MH428965 and MH428966, results showed that the phytoplasmas from group 16SrIX detected in periwinkle and Gliricidia in Costa Rica are different from each other, and the strain found in periwinkle may represent a variant of subgroup 16SrIX-A, meanwhile Costa Rican GLLD strains have 100 % identity to those from Honduran GLLD.
The presence of group 16SrXIII, ‘Ca. Phytoplasma hispanicum’, in Costa Rica was first detected in this survey. Two plants showing virescence and harboring this phytoplasma group were collected in Dulce Nombre (Cartago) and Hojancha (Guanacaste, GenBank accession no. MH428958). The 16S rRNA gene sequences obtained and compared by the BLAST algorithm showed 99 % identity with Mexican periwinkle virescence phytoplasma (MPVP, GenBank accession no. AF248960). According to virtual RFLP analysis, it was most similar to the reference pattern of subgroup 16SrXIII-H (GenBank accession no. JX626329), with a similarity coefficient of 0.98, suggesting this may be a variant of 16SrXIII-H. The virtual RFLPs displayed patterns like those of subgroup 16SrXIII-H, except for a unique pattern for enzyme MseI (Figure 3D).
The fifth phytoplasmas group detected during this work was ‘Ca. Phytoplasma brasiliense’ (16SrXV). Infection of C. roseus by this group was confined to Guanacaste province. All three plants harboring group 16SrXV showed virescence; with one plant (CR03) collected in Samara (Guanacaste) also exhibiting phyllody and another one (CR14, Santa Cruz, Guanacaste) also exhibiting big bud (Figure 1E). The 16S rRNA gene sequences obtained showed 99 % similarity with sequences corresponding to this phytoplasma infecting Vitis vinifera, Carica papaya, Crotalaria juncea, Hibiscus sp. and Guazuma ulmifolia from Peru (GenBank accession nos. KX670807-9, KX810334-6), Brazil (GenBank accession nos. KF878382 and AF147708), and Costa Rica (GenBank accession nos. HQ258882-3), respectively. However, an in silico RFLP comparison using the iPhyClassifier showed that the phytoplasma found in C. roseus is not the same as the 16SrXV-B subgroup previously reported in Costa Rica infecting G. ulmifolia. According to the virtual RFLP pattern obtained for the sequence of GenBank accession no. MH428963, the most similar to it was the reference pattern of subgroup 16SrXV-A (GenBank accession no. AF147708), with a similarity coefficient of 0.96, suggesting this strain may represent a tentative new 16SrXV subgroup. The virtual RFLP analysis displayed a unique and different RFLP pattern with HaeIII to previously reported strains (Figure 3E). The 16 remaining virtual RFLPs showed exactly the same patterns to those of subgroup 16SrXV-A, for example with enzyme HpaII (Figure 3F).
Phylogenetic analysis comparison based on 16S rDNA sequences of phytoplasma subgroups found in this study showed 16Sr group clustering consistent with the group identification by BLAST analysis. The same consistent phylogenetic clustering was found with representatives of groups 16SrIII, 16SrIX and 16SrXIII. However, representatives of subgroups 16SrI-B and 16SrI-P, obtained in this study, clustered together instead of with related sequences obtained from GenBank, suggesting a possible relationship due to geographic origin. Also, contrary to the iPhyClassifier results, the representative strain of group 16SrXV from C. roseus, obtained in this study, clustered more closely to subgroup 16SrXV-B strains than subgroup 16SrXV-A (Figure 4).
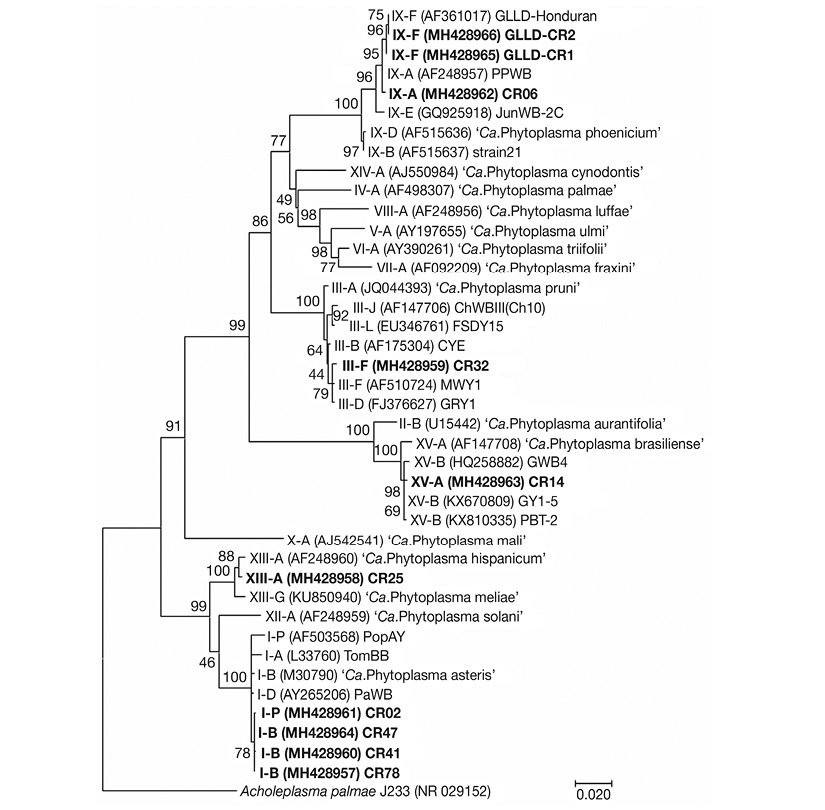
Figure 4 Phylogenetic tree inferred from analysis of 16S rRNA gene sequences by Maximum Likelihood method based on the General Time Reversible model. A bootstrap test with 2500 replicates was done to support reliability of the analysis. The analysis involved 41 nucleotide sequences, included reference strains of 30 previously described Ca. Phytoplasma species and subgroup strains obtained from GenBank and Acholeplasma palmae J233 (NR_029152), as outgroup to root the phylogenetic tree. GenBank accession numbers of all sequences are showed in parenthesis and sequences obtained in this study are indicated in bold font. Bar = 0.02 nucleotide substitutions per site.
Discussion
In this study, phytoplasmas were detected in naturally infected C. roseus, in six of seven provinces in Costa Rica (excluding Alajuela), 16SrRNA gene sequences indicated that they belonged to five groups: 16SrI, 16SrIII, 16SrIX, 16SrXV, and 16SrXIII. The first four groups were previously identified in other plant hosts in Costa Rica (Kenyon et al., 1999; Villalobos et al., 2002; 2011; Pardo et al., 2014). Group 16SrXIII was detected in the country for the first time. Moreover, three new tentative subgroups into 16SrI, 16SrIX and 16SrXV groups were also described. These findings suggested that the phytoplasmas present in Costa Rica are more diverse and complex than previously known, and these rise potential risk of phytoplasmal diseases to economically crops as well as wild biodiversity. Despite the presence of two different phytoplasmas in some locations in the same province (Table 2), no mixed infections were found in this study probably because few clones per plant were analyzed.
Catharanthus roseus has been reported as a natural host for phytoplasmas groups 16SrI, 16SrIII, 16SrIX, 16SrXV, 16SrXIII, in North and South America, and the Caribbean islands, but not from the Central American region (Lee et al., 1998b; Lee, Davis, & Gundersen-Rindall, 2000; Montano, Dally, Davis, Pimentel, & Brioso, 2001b; Torres, Galdeano, Docampo, & Conci, 2004; Duduk, Mejia, Calari, & Bertaccini, 2008; Barbosa, Eckstein, Bergamin Filho, Bedendo, & Kitajima, 2012; Galdeano, Guzmán, Fernández, & Conci, 2013; Dumonceaux, Green, Hammond, Pérez, & Olivier, 2014; Caicedo, Rivera-Vargas, Segarra, & Davis, 2015; Davis, Harrison, Zhao, Wei, & Dally, 2016; Pérez-López, Luna-Rodríguez, Olivier, & Dumonceaux, 2016a). According to Montano et al. (2001b), C. roseus is a well-known experimental host for phytoplasmas, however few incidences of natural infections have been reported and its role related to natural dissemination and disease spread is uncertain. Nevertheless, considering a different perspective, C. roseus may be deployed as a sentinel plant for phytoplasma occurrence in a geographical area, as this study is showing.
Subgroup 16SrI-B phytoplasma strains (‘Ca. Phytoplasma asteris’) have been detected in Costa Rica associated with diseases affecting crops, landscape trees, and wild shrubs (Gámez & León, 1985; Villalobos et al., 2002, Saborío-Rodríguez, Villalobos, & Rivera, 2007). Two epidemics associated with this phytoplasma subgroup caused severe economic losses to chayote (Sechium edule) growers during 2001-2002, and to coffee growers during 2006-2007 due the infection of Erythrina trees, which are used as shade trees for coffee crop production in Costa Rica (Saborío-Rodríguez et al., 2007). In this study, this subgroup was also the most widespread in the country. We hypothesize that 16SrI group has been present longer in the country; therefore, allowing the emergence of new host-vector interactions, recently three wild plant species belonging to Anacardiaceae (Villalobos, unpublished), Asteraceae / Compositae and Rubiaceae (Villalobos, Montero-Astúa, Coto, Sandoval, & Moreira, 2018) have been detected as new hosts for this group.
The phytoplasma strain variant of subgroup 16SrI-P found in Turrialba (Cartago province, GenBank accession no. MH428961) has not been previously reported in Costa Rica or Central America. This subgroup was reported by Šeruga et al. (2003) infecting Populus nigra trees in Croatia. However, BLASTn top hit results showed 99 % similarity of the Turrialba’s periwinkle sample with other phytoplasmas found in North America, including a periwinkle with virescence collected in Yucatan, Mexico (GenBank accession no. EF050085).
Likewise, the detection of phytoplasmas belonging to the 16SrIII-F subgroup (‘Ca. Phytoplasma pruni’ related strain) is a new report for Costa Rica. This subgroup was also detected in a different host in Cartago province (Villalobos, unpublished data). Pardo et al. (2014) reported subgroup 16SrIII-L infecting cassava crop associated with frog-skin disease in the country. The 16SrIII group has a large number of subgroups and diversity of host species, as well as a wide geographic distribution (Zhao et al., 2009). For example, subgroup 16SrIII-J is notable for its presence in several hosts and several countries mainly in South America (Galdeano et al., 2013, Pérez-López et al., 2016a).
It is highlighted that natural infections with 16SrIX-A are reported for the first time in Costa Rica. Its presence in at least two geographically distant sites, Southern area (Potrero Grande, Puntarenas) and Central Valley (San Pedro, San José), may suggest a high probability of being found in other areas. The symptoms in both geographic locations are consistent with those previously reported for the host and subgroup 16SrIX-A, such as virescence and yellowing. Natural infection of this plant species by group 16SrIX-A was reported previously in Colombia (Duduk et al., 2008), Brazil (Barbosa et al., 2012) and Puerto Rico (Caicedo et al., 2015).
Reports of phytoplasmas belonging to group 16SrXIII have been concentrated in relatively few hosts and restricted geographic locations inside North and South America, infecting C. roseus, Solanum tuberosum, C. papaya, S. lycopersicum, Fragaria sp., Melia azedarach, Turnera ulmifolia, Dimorphandra spp. and Thunbergia erecta (Lee et al., 1998b; Harrison, Boa, & Carpio, 2003; Arneado et al., 2007; Montano et al., 2011; Melo et al., 2013; 2018; Montano, Bertaccini, Guthelle, Paltrinieri, & Contaldo, 2015; Fernández et al., 2015; Alves, Souza, Ribeiro, da Silva Xavier, & Carvalho, 2016). The findings reported in this study, expand the geographical range of ‘Ca. Phytoplasma hispanicum’ (Davis et al., 2016). In this survey, plants infected with subgroup 16SrXIII-A were collected in Cartago province and Guanacaste, and no other hosts have been detected in the country for this phytoplasma group. Additional analyses must be done to determine if this phytoplasma may represent a new subgroup in accordance with differential virtual RFLP patterns, as mentioned above.
Phytoplasmas of group 16SrXV, ‘Ca. Phytoplasma brasiliense’, were reported in Brazil infecting Hibiscus spp. (Montano et al., 2001a), and other hosts including C. roseus, Crotalaria juncea Brassica oleracea, and Sida rhombifolia (Montano et al., 2001b; Eckstein, Barboza, Rezende, & Bedendo, 2011; Bianco, et al., 2014; Canale, & Bedendo, 2013). It is also present in Costa Rica infecting Guazuma ulmifolia (Villalobos, et al., 2011), and in Peru infecting Vitis vinifera and C. papaya (Wei, et al., 2017). The Costa Rican Guazuma strain was classified as subgroup 16SrXV-B and the Peruvian strain was identified as a variant. The vinca samples infected with group 16SrXV were found in different places in the Nicoya peninsula (Guanacaste), where the presence of Guazuma witches’-broom (GWB) is frequent. Nevertheless, sequences showed 99 % sequence similarity to ‘Ca. Phytoplasma brasiliense’ representatives of subgroup 16SrXV-B variants from Peru (GenBank accession nos. KX670807-9, KX810334-6), rather than to the surrounding Guazuma phytoplasma. Symptoms associated to natural infection of C. roseus by this phytoplasmas in Brazil (Montano et al., 2001b) were yellowing and witches’-broom, while in Costa Rica the main symptom was virescence.
As previously reported by Gundersen et al. (1996b) and Lee et al. (1998a, 2004), genetic variation in some phytoplasma strains appears to be associated with ecological isolation, and distinct phytoplasma strains may result from new epidemiological cycles from an original organism (Lee et al., 1998b). The tentative variants and or tentative subgroups observed in phytoplasmas sequenced in this study may be the result of ecological isolation in the C. roseus host. It is worth noting that we observed in this study a case where the host species and/or the putative insect vector associated with the plant host may represent the ecological differentiator for a phytoplasma strain, rather than geographic relationship. Catharanthus roseus plants with phytoplasma infection were collected from Guanacaste, where Guazuma trees with subgroup 16SrXV-B phytoplasma infection are common; nevertheless, contrary to our preliminary hypothesis, the C. roseus plant growing in Guanacaste, in the same area where Guazuma symptomatic trees occur, were infected with different phytoplasma strains.
Including the results from this report, phytoplasmas belonging to six groups and nine subgroups have been detected in Costa Rica. Moreover, phytoplasma infection has been detected in more than 17 different plant hosts. These findings suggest more diversity of phytoplasmas in the country than previously suspected. Attention to their incidence and geographic spread is important because we hypothesize that new host encounters are allowing the development of previously uncharacterized host-phytoplasma associations. On the other hand, accidental or intentional introduction of phytoplasmas to a new geographical region must be avoided due to the effects that these pathogens may cause to biodiversity. Examples of this situation increase every year in scientific reports about new diseases, new hosts, or new geographic range involving these plant pathogens.
This study extends the knowledge of phytoplasmas subgroups in Costa Rica and the Central American region. Here we report for the first time the detection of natural infection of C. roseus with subgroups 16SrI-B, 16SrI-P, 16SrIII-F, 16SrIX-A, 16SrXIII-A, 16SrXV-A. With the exception of 16SrI-B, all of these subgroups were previously unknown to occur in the Central American region. The findings here reported are not only informative, these identify C. roseus as a reservoir and potential inoculum source of phytoplasmas to cultivated crops, wild plants, as well as other ornamental plants. Therefore, this plant species may be considered a sentinel to detect phytoplasmas within surrounding crops and natural ecosystems, as well as their genetic diversity.
Ethical statement: authors declare that they all agree with this publication and made significant contributions; that there is no conflict of interest of any kind; and that we followed all pertinent ethical and legal procedures and requirements. A signed document has been filed in the journal archives












 uBio
uBio 

