Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO -
 Access statistics
Access statistics
Related links
-
 Similars in
SciELO
Similars in
SciELO  uBio
uBio
Share
Revista de Biología Tropical
On-line version ISSN 0034-7744Print version ISSN 0034-7744
Rev. biol. trop vol.62 n.4 San José Oct./Dec. 2014
Development of RAPD-SCAR markers for Lonicera japonica (Caprifolicaceae) variety authentication by improved RAPD and DNA cloning
Desarrollo de marcadores RAPD-SCAR para Lonicera japonica (Caprifolicaceae) para la autenticación de variedades mediante RAPD mejorados y clonación de ADN
Desarrollo de marcadores RAPD-SCAR para Lonicera japonica (Caprifolicaceae) para la autenticación de variedades mediante RAPD mejorados y clonación de ADN
Luquan Yang1*, Md. Asaduzzaman Khan1, Zhiqiang Mei1, Manman Yang1, Tiandan Zhang1, Chunli Wei1, Weichan Yang1, Li Zhu1, Yan Long1 & Junjiang Fu1,2*
Abstract
Genetic diversity within a species is a common feature, which plays a vital role in its survival and adaptability, and is important for the identification and authentication of a species. Lonicera japonica is a traditionally used medicinal plant, which have been recently genetically characterized by an improved ran- dom amplified polymorphic DNA (RAPD) analysis. In this study, the molecular markers on the basis of these RAPD fragments have been developed to identify specific L. japonica variety. The DNAs were extracted from fresh young leaves of different samples of L. japonica collected from Shenzhen, Yichang, Leshan, Emei and Loudi, China. The DNA materials were amplified using improved RAPD PCR. Different RAPD bands were excised, cloned and developed for stable sequence-characterized amplified region (SCAR) markers with different species. Two SCAR markers, JYH3-3 and JYH4-3, have been successfully cloned from improved RAPD fragments. The SCAR marker JYH3-3 was found specific for all of the L. japonica samples collected from the different regions, and another marker JYH 4-3 was strictly specific to the Shenzhen sample from Guangdong province, which is geographically distant from Hubei, Sichuan and Hunan Provinces (source of other L. japonica samples). The marker JYH3-3 was found as specific molecular marker for the identification of L. japonica, while JYH4-3 was found as molecular marker strictly specific for the Shenzhen sample. The developed SCAR mark- ers might serve as more specific molecular markers for L. japonica variety authentication. The combination of improved RAPD analysis and SCAR marker development have resulted useful tools to study the genetic variety of any organism, which we have successfully applied here in L. japonica.
Key words: molecular markers, Lonicera japonica, random amplified polymorphic DNA, sequence-characterized amplified region, identification of genetic species.
Resumen
La diversidad genética dentro de una especie es una característica común, que juega un papel vital en su supervivencia y adaptabilidad, y es importante para la identificación y la autenticación de una especie. Lonicera japonica es una planta medicinal utilizada tradicionalmente, que han sido recientemente caracterizada genéticamente por amplificación aleatoria mejorada de ADN polimórfico (RAPD). En este estudio, los marcadores moleculares basados en estos fragmentos de RAPD se han desarrollado para identificar una variedad específica de L. japonica. Los ADN se extrajeron de las hojas jóvenes frescas de diferentes muestras de L. japonica recogidas de Shenzhen, Yichang, Leshan, Emei y Loudi, China. Los materiales de ADN fueron amplificados utilizando el RAPD PCR mejorado. Diferentes bandas RAPD fueron extraídas, clonadas y desarrolladas para las regiones amplificadas de secuencia conocida (SCAR) con marcado- res de diferentes especies. Dos marcadores SCAR, JYH3-3 y JYH4-3, se clonaron con éxito de los RAPD mejorados. El marcador SCAR JYH3-3 se encontró específico para todas las muestras de L. japonica recolectadas en las diferentes regiones, mientras que el otro marcador JYH4-3 era estrictamente específico para la muestra de Shenzhen de la provincia de Guangdong, que está geográficamente distante de Hubei, Sichuan y Provincias Hunan (fuente de otras muestras de L. japonica). Se encontró que JYH3-3 es un marcador molecular específico para la identificación de L. japonica, mientras que JYH4-3 se encontró como marcador molecular estrictamente específico para la muestra de Shenzhen. Los marcadores SCAR desarrollados podrían servir como marcadores moleculares más específicos para la autenticación de la variedad L. japonica. La combi- nación de RAPD mejorado y el desarrollo del marcador SCAR han dado como resultado herramientas útiles para el estudio de la variedad genética de cualquier organismo, que hemos aplicado con éxito en L. japonica.
Palabras clave: marcadores moleculares, Lonicera japonica, ADN polimórfico amplificado al azar, regiones amplificadas de secuencia conocida, identificación de especies genéticas.
Genetic differences within a species are a common feature in the living world, and a result of the adaptation to changing environments. It has been postulated that genetic diversity plays a vital role in the survival and adaptability of a species, and its escape from the extinction with changing environmental conditions (Frankham, 2008). Besides, the analysis of the genetic diversity is important for the identification and authentication of a species; it is also important for organisms genetic profiling and conservation. Considering the analysis of the genetic diversity, a number of molecular marker techniques have been developed over the last 30 years: random amplified polymor- phic DNA (RAPD), simple sequence repeat (SSR), inter-simple sequence repeat (ISSR) and amplified fragment length polymorphism (AFLP) analysis (Agarwal, Shrivastava, & Padh, 2008; Liu, Li, Khan, & Zhu, 2012; Fu, Yang, Khan, & Mei, 2013). These techniques are currently being used for the genetic characterization and identification of different unicel- lular and multicellular organisms.
Lonicera japonica Thunb., known as the Japanese honeysuckle, or Jin-Yin-Hua (JYH) in Chinese, is a traditional medicine used mainly in some parts of East-Asian countries, including China, Japan and Korea. L. japonica is well known for its anti-cancer, anti-inflammatory, anti-virus, anti-angiogenic, anti-oxidant, hepato-protective and wound healing activities (Xiang et al., 2001; Zhang, Song, & Shi, 2011; Chen, Liou, Tzeng, Lee, & Liu, 2012; Park et al., 2012, Cho et al., 2012). Although not under the extinction threat, this medicinal plant is not widely recognized all over the world. One main reason behind the narrow spectrum regional use of this plant, is the lack of genetic information and invalidated identification process. Recently, by employing improved RAPD analysis, we have genetically characterized and authenticated L. japonica species from differ- ent regions of China, which are geographically isolated (Fu et al., 2013). Sequence-character- ized amplified region (SCAR) marker is one of the stable molecular markers that is generally derived from RAPD, of which the basic principle is to convert the dominant markers into co-dominant markers to reduce the tedious- ness of RAPD (Dnyaneshwar, Preeti, Kalpana, & Bhushan, 2006; Li, Tang, & Cai, , 2010; Rajesh et al., 2013). SCAR markers usually reveal higher levels of polymorphism owing to higher annealing temperatures and longer primer sequence specificity (Kumla, Doolgin- dachbaporn, Sudmoon, & Sattayasai, 2012). When RAPD is combined with SCAR markers, the analytical procedure becomes a simple PCR analysis, using PCR primers designed from the sequence of RAPD amplicons (Kumla et al., 2012; Rajesh et al. 2013), as we have indicated earlier (Fu et al., 2013). In this study, we have developed SCAR markers from previously established RAPD fragments for the genetic characterization, authentication and validation of L. japonica.
Materials and methods
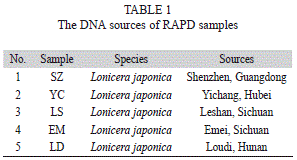
Extraction of L. japonica DNA: The DNAs were extracted from fresh young leaves of different samples of L. japonica (Table 1) by using previously described method, diluted with a final concentration of 10ng/μL and stored at -20°C until use (Fu et al., 2013).
Amplification of DNA by improved RAPD: The improved RAPD PCR were ini- tially amplified with random primers SBC- Q2 and SBC-I10 using above mentioned L. japonica DNAs with Tiangen reagents (Beijing, China). A total 15μL PCR reaction system was consisted of 7.5μL 2×Taq PCR MasterMix, 1.5μL 2.5μM primer, 1.5μL genomic DNA, and with ddH2O. Amplification reactions were per- formed by using “Applied Biosystems Veriti® 96-Well Thermal Cycler” (Life Technology, USA), with the following steps: initial denaturation at 95°C for 90s, 40 cycles of denaturation at 94°C for 40s, annealing at 36°C with the RAMP rate from annealing to extension was adjusted to 0.125ºC/s (5% ramp rate) for 60s, extension at 72°C for 90s, and a final extension step at 72°C for 5min. PCR products were detected with 1.5% agarose gel electrophoresis.
Cloning and sequencing of DNA fragments: Two different bright bands were excised from agarose gel, purified by using TIANgel Mini Purification Kit (DP209, China). Puri- fied DNA fragments were ligated into pGM-T vector (No. VT202, Tiangen reagents, Beijing, China), and transformed into DH5α E. coli competent cells. The recombinant clones were selected on LB agar plates containing 100μg/ μL of ampicillin, 40mg of X-gal and 160μg of IPTG. The white colonies were screened out by blue white screening. The presence of right insert was verified by PCR by using T7/SP6 primer pairs (T7 primer: 5’-TAATACGACT- CACTATAGGG-3’, SP6 primer: 5’-ATTTAG- GTGACACTATAGAA-3’), which is located at pGM-T vector near to the ligation ends, and EcoRI digestion (Fu, 2012). The cloned DNA fragments were then sequenced by Sanger method. The homology of sequenced DNA was searched and analyzed by the online pro- gram BLAST (http://www.ncbi.nlm.nih.gov/ BLAST/) in different species.
SCAR primer design: The nucleotide sequence of each of the cloned RAPD fragment was used to design pairs of SCAR primers using Primer 3 (http://bioinfo.ut.ee/ primer3-0.4.0/primer3/) and sequences of each primer and amplification length were listed in table 2.
Development SCAR markers and SCAR analysis: To develop SCAR markers, the PCR amplification was performed, where the content of 10μL PCR reaction system was as follows: 5μL 2×Taq PCR MasterMix, 1μL of 2.5μM each pair of SCAR primers and 1μL genomic DNA (10ng), the remaining volumes were filled by ddH2O. PCR was performed by using “Applied Biosystems Veriti® 96-Well Thermal Cycler” (Life Technology, USA) with an initial pre-denaturation for 90s at 95°C, followed by 35 cycles of denaturation at 94°C for 40s, annealing at different temperatures 54°C, 57°C, 60°C, 63°C or 66°C for 30s, and extension at 72°C for 40s. The final exten- sion step was performed at 72°C for 5 min. The amplified PCR products were separated by electrophoresis on 1.0% agarose gel in 1×TAE buffer. Gels were visualized by 0.5μg/ mL ethidium bromide staining, and the images were documented using the ChemiDoc XRS (Bio-Rad, USA). Sequences of the SCAR primers, amplified length and PCR condition were included in table 2.
To distinguish the difference between the varieties of L. japonica and other species, the SCAR analysis was performed by using 20 DNA samples as templates, including five samples of L. japonica which were described previously (Fu et al., 2013), five samples of D. longan collected from Sichuan, Guang- dong, Guangxi, Fujian and Hainan (Yang, Fu, Khan, Zeng, & Fu 2013; Mei et al., 2014), five samples of C. album collected from Luzhou City, Gastrodia elata collected from Liangshan City in Sichuan Province (Mei et al., 2014), P. chinese collected from Gulin County in Sich- uan province, P. sedoides, D. confinis collected from Guangxi source of wild, and V. philippica (Yang et al., 2013). PCR amplifications were performed by using above mentioned 2 pairs of SCAR primers and amplification conditions in table 2.
Results
Cloning of RAPD fragments: Two RAPD primers, SBC-Q2 (Q2) and SBC-I10 (I10), were used to improve RAPD amplification from five DNA samples of L. japonica, which were collected from Shenzhen City of Guangdong Province, Yichang City of Hubei Province, Leshan and Emei City of Sichuan Province and Loudi City of Hunan Province, respectively (Fu et al., 2013). The results are shown in the figure 1, the black arrows indi- cated bands labeled with JYH3 in primer Q2 (Fig. 1A) and labeled with JYH4 in primer I10 (Fig. 1B). The DNA bands with black arrows were cut from the agarose gel, purified, ligated to T-vector by AT cloning, and under black and white screening method. The positive clones were then identified by plasmid DNA digestion using EcoRI enzyme in clones JYH 3-3 (Fig. 1C) and JYH 4-3 (Fig. 1C), or by PCR amplification using SP6 primer and T7 primer in clone JYH 4-3 (Fig. 1E). In the figure 1C, clone JYH 3-3, which showed with a similar ~600-700bp inserted DNA-fragment, was sequenced. In the figure 1E, clones JYH4-1, JYH4-2, JYH4-3 showed PCR band with a similar ~300bp inserted DNA-fragment, and clone JYH4-3 was verified by EcoR I digestion (Fig. 1D), and was selected for further sequencing.
Sequences and characterization of L. japonica-specific RAPD fragments: Sequencing of the above two cloned RAPD fragments of L. japonica revealed that the clone JYH4-3 consisted of 589 nucleotides and was deposited into GenBank with accession number KF698799 (Fig. 2A), and the clone JYH4-3 consisted of 308 nucleotides and was deposited into GenBank with accession number KF698800 (Fig. 2B). BLAST searches of the nucleotide sequences in GenBank showed that 458 nucleotides of clone JYH3-3 fragment (nucleotides 8 to 588) shared 78% identity to the mRNA of Vitis vinifera probable receptor-like protein kinase At1g67000-like (LOC100253816) (Sequence ID: ref|XM_002268656.2|) with an E value 4e-97 (Fig. 2C). The nucleotide sequences of clone JYH 4-3 fragment did not show any identity to that of any species (data not shown).
Development of L. japonica-specific SCAR markers JYH3-3 and JYH4-3: To generate a stable L. japonica-specific diagnostic SCAR markers from RAPD markers, two pairs of primers (JYH 3-3L and JYH 3-3R; JYH 4-3L and JYH 4-3R) (Table 2) were designed and synthesized based on cloned sequences in figure 2. The PCR amplification results with different annealing temperature are shown in figure 3 A, figure 3 B, which indicates that annealing with 63°C have better specific amplification in SCAR maker JYH3-3 with expected size (Fig. 3 A), and with 60°C have better amplification in SCAR maker JYH4-3 with expected size in the L. japonica samples (Fig. 3 B). Negative control (NC) without DNA template did not show any PCR products. Therefore, L. japonica-specific SCAR markers were developed. The sequences of SCAR primers, amplified length and PCR condition are shown in table 2.
Authentication L. japonica: The designed SCAR primer pairs were then used to amplify the genomic DNA from 20 of collected DNA samples to test amplification species-specificity. The PCR amplification was performed again. The results indicated that the PCR products with expected size were observed in all five of L. japonica samples by SCAR marker JYH3-3, without any amplification in other species we tested (Fig. 3C). This indicates that SCAR marker JYH3-3 is L. japonica-specific. The lack of this specific amplicon in the other species indicates the efficacy of this marker in distinguishing the L. japonica group from the others. The PCR results by SCAR marker JYH4-3 indicated that the PCR products with expected size were observed only in Shenzhen sample of L. japonica, without any amplification in other four L. japonica samples and in other species we tested (Fig. 3D). This indicated that the SCAR marker JYH4-3 is Shenzhen sample-specific. The lack of this specific amplicon in the samples from Yichang, Leshan, Emei and Loudi city and other species indicates the efficacy of this marker in distinguishing the Shenzhen sample of L. japonica from other cultivars, as well as other species.
Discussion
In modern bio-molecular science, molecular marker technologies have become significant tools. These technologies are helpful for systematic biologists, and very useful to solve species and population identification level problems. RAPD is one of the frontline tech- niques, which is widely used in genetic characterization and identification of any organism. It is easy to handle; it can reveal high degrees of polymorphisms; and it does not require prior DNA sequence information of the species (Williams, Kubelik, Livak, Rafalski, & Tingey, 1990; Shakeel, Ilyas & Kazi, 2013; Noormo-hammadi, Hasheminejad-Ahangarani, Sheidai, Ghasemzadeh-Baraki, & Alishah, 2013; Fu et al., 2013; Kumar et al., 2013). In recent years, a number of plants and other organisms have been characterized genetically by RAPD and improved RAPD analysis (Kumla et al., 2012; Bhat et al., 2012, Fu et al., 2013). Moreover the potential of RAPD has also been proven in the study of genetic diversity in newly found or synthetic species of organisms, which are important both in agriculture and industry (Shakeel et al., 2013). It has been established that conversion of RAPD markers into SCAR markers can improve the specificity and stability, which makes this technique more convenient and efficient in the testing of different alleles (Rajesh et al., 2013). As SCAR markers can identify a single or a few bands instead of a complex pattern, they are more straightforward than RAPD, SSR, ISSR and AFLP. So identification of any organism becomes more authentic and well-verified if RAPD analysis is combined to SCAR marker technology.
The recent resurgence of traditional medicine research has significant importance in modern biomedical research. For this, identification and characterization of medicinal plants is of emerging importance among the group of biologists. L. japonica is a well-known Chinese and Japanese traditional medicinal plant. In a recent study, an improved RAPD analysis have reported the genetic distance between the samples of this plant collected from different geographic locations of China (Fu et al., 2013). Here in this study, we have generated SCAR markers from the cloning of previously developed RAPD markers (Fu et al., 2013). For this we have also optimized the anneal- ing temperature, and we found that at 60oC, or 63oC, amplification was better than other that in other temperatures (Table 2). This study reports that the marker JYH3-3 is specific to the DNAs from all L. japonica samples, as they showed no PCR amplification in DNA of other plants used in this study (D. longan, C. album, Gastrodia elata, P. chinese, P. sedoides, D. confinis, V. philippica). Interestingly, another marker JYH4-3 was found to be Shenzhen sample-specific. This finding reflects our previous study (Fu et al., 2013), where we reported that Shenzhen is geographically far-isolated from other regions, and possess high level of genetic distance. This study actually validates the improved RAPD analysis result of our previous study, as well as establishes specific molecular markers (SCAR) to distinguish L. japonica from other plants, and even to differ- entiate within species. According to the best of our knowledge, this is the first RAPD-SCAR study on L. japonica identification. However, some studies have reported the RAPD-SCAR technique for the identification and authentication of other organisms like pepper (Jiang, Jianhua, & Li, 2009), longan fruits (Yang et al., 2013), shrimp (Dutta et al., 2014), silkworm (Dutta et al., 2012), microbes (Lee, Lee, Lee & Lee, 2013) among others.
This study sucessfully developed specific SCAR markers for the identification of the medicinal plants of L. japonica. This study also indicated that improved RAPD analysis is a potential molecular technique for the genetic characterization and identification of any species, and when the RAPD fragments are cloned and generated to SCAR markers, they can serve as more specific molecular markers. Thus, combining these two molecular marker techniques can provide a simple and reliable tool for the genetic characterization, identification, authentication of other medicinal plants as well as any other plant species.
Acknowledgments
This research was supported in part by the National Natural Science Foundation of China (30371493 and 81172049), the Science and Technology Innovation Team of Colleges and Universities in Sichuan Province (13TD0032), and the Health Department Foundation of Sichuan Province (130261). The authors particularly thank all individuals who provided DNAs or leaves for L. japonica and other species.
References
Agarwal, M., Shrivastava, N., & Padh, H. (2008). Advances in molecular marker techniques and their appli- cations in plant sciences. Plant Cell Reports, 27(4), 617-631. [ Links ]
Bhat, A. A., Haniffa, M. A., Divya, P. R., Gopalakrishnan, A., Milton, M. J., Kumar, R., & Paray, B. A. (2012). Molecular characterization of eight Indian Snake-head species (Pisces: Perciformes Channidae) using RAPD markers. Molecular Biology Reports, 39(4), 4267-4273. [ Links ]
Chen, W. C., Liou, S. S., Tzeng, T. F., Lee, S. L., & Liu, I. M. (2012). Wound repair and anti-inflammatory potential of Lonicera japonica in excision wound-induced rats. BMC Complementary and Alternative Medicine, 12, 226. [ Links ]
Cho, W. K., Kim, H., Choi, Y. J., Yim, N. H., Yang, H. J., & Ma, J. Y. (2012). Epimedium koreanum Nakai water extract exhibits antiviral activity against porcine epidermic diarrhea virus in vitro and in vivo. Evidence Based Complementary and Alternative Medicine, 2012, 985151. [ Links ]
Dnyaneshwar, W., Preeti, C., Kalpana, J., & Bhushan, P. (2006). Development and application of RAPD- SCAR marker for identification of Phyllanthus emblica LINN. Biological & Pharmaceutical Bulletin, 29(11), 2313-2316. [ Links ]
Dutta, S. R., Kar, P. K., Srivastava, A. K., Sinha, M. K., Shankar, J., & Ghosh, A. K. (2012). Identification of RAPD and SCAR markers associated with yield traits in the Indian tropical tasar silkworm Antheraea mylitta drury. Genetics and Molecular Biology, 35(4), 743-751. [ Links ]
Dutta, S., Biswas, S., Mukherjee, K., Chakrabarty, U., Mallik, A., & Mandal, N. (2014). Identification of RAPD-SCAR marker linked to white spot syndrome virus resistance in populations of giant black tiger shrimp, Penaeus monodon Fabricius. Journal of Fish Diseases, 37(5), 471-480. doi: 10.1111/jfd.12128. [ Links ]
Frankham, R. (2008). Genetic adaptation to captivity in species conservation programs. Molecular Ecology, 17(1), 325-333. [ Links ]
Fu, J. J. (2012). Short protocols in medical molecular biology. Beijing: China Medical Science Press. [ Links ]
Fu, J., Yang, L., Khan, M. A., & Mei, Z. (2013). Genetic characterization and authentication of Lonicera japonica Thunb. by using improved RAPD analysis. Molecular Biology Reports, 40(10), 5993-5999. [ Links ]
Jiang, S., Jianhua, X., & Li, X. (2009). A Study on the RAPD and SCAR Molecular Markers of Piper Species. Journal of Agriculture and Rural Development in the Tropics and Subtropics, 110(2), 127-135. [ Links ]
Kumar, S., Sharma, R., Kumar, V., Govind, K., Vyas, G., & Rathore, A. (2013).Combining molecular-marker and chemical analysis of Capparis decidua (Capparaceae) in the Thar Desert of Western Rajasthan (India). Revista de Biología Tropical, 61(1), 311-320. [ Links ]
Kumla, S., Doolgindachbaporn, S., Sudmoon, R., & Sattayasai, N. (2012). Genetic variation, population structure and identification of yellow catfish, Mystus nemurus (C&V) in Thailand using RAPD, ISSR and SCAR marker. Molecular Biology Reports, 39(5), 5201-5210. [ Links ]
Lee, D. H., Lee, S. K., Lee, S. Y., & Lee, J. K. (2013). Development of SCAR Markers for the Identification of Phytophthora katsurae Causing Chestnut Ink Disease in Korea. Mycobiology, 41(2), 86-93. [ Links ]
Li, S. F., Tang, S. J., & Cai, W. Q. (2010). RAPD- SCAR markers for genetically improved NEW GIFT Nile Tilapia (Oreochrmis niloticus niloticus L.) and their application in strain identification. Zoological Research, 31(2), 147-153. [ Links ]
Liu, W., Li, L., Khan, M. A., & Zhu, F. (2012). Popular molecular markers in bacteria. Molecular Genetics, Microbiology & Virology, 27(3), 103-107. [ Links ]
Mei, Z., Fu, S., Yu, H., Yang L., Duan, C., Liu, Y., Gong, S., & Fu, J. (2014). Genetic characterization and authentication of Dimocarpus longan Lour. using an improved RAPD technique. Genetics and Molecular Research, 13(1), 1447-1455. [ Links ]
Mei, Z., Yang, L., Zhang, T., Yu, H., Gan, L., Zhang, L., Yang, M., & Fu, J. (2014). Improved RAPD Analysis of Canarium album (Lour.) Raeusch from Sichuan Province along Yangtze River in China. Annual Research & Review in Biology, 4(1), 51-60. [ Links ]
Noormohammadi, Z., Hasheminejad-Ahangarani, F. Y., Sheidai, M., Ghasemzadeh-Baraki, A. S., & Alishah, O. (2013). Genetic diversity analysis in Opal cotton hybrids based on SSR, ISSR, and RAPD markers. Genetics and Molecular Research, 12(1), 256-269. [ Links ]
Park, H. S., Park, K. I., Lee, D. H., Kang, S. R., Nagappan, A., Kim, J. A., Kim, E. H., Lee, W. S., Shin, S. C., Hah, Y. S., & Kim, G. S. (2012). Polyphenolic extract isolated from Korean Lonicera japonica Thunb. Induce G2/M cell cycle arrest and apoptosis in HepG2 cells: involvements of PI3 K/Akt and MAPKs. Food and Chemical Toxicology, 50(7), 2407-2416. [ Links ]
Rajesh, M. K., Jerard, B. A., Preethi, P., Thomas, R. J., Fayas, T. P., Rachana, K. E., & Karun, A. (2013). Development of a RAPD-derived SCAR marker associated with tall-type palm trait in coconut. Scientia Horticulturae, 150, 312-316. [ Links ]
Shakeel, M., Ilyas, M., & Kazi, M. (2013). Evaluation of synthetic hexaploid wheats (derivative of durum wheats and Aegilops tauschii accessions) for studying genetic diversity using randomly amplified polymorphic DNA (RAPD) markers. Molecular Biology Reports, 40(1), 21-26. [ Links ]
Williams, J. G., Kubelik, A. R., Livak, K. J., Rafalski, J. A., & Tingey, S. V. (1990). DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Research, 18(22), 6531-6535. [ Links ]
Xiang, T., Xiong, Q. B., Ketut, A. I., Tezuka, Y., Nagaoka, T., Wu, L. J., & Kadota, S. (2001). Studies on the hepatocyte protective activity and the structure- activity relationships of quinic acid and caffeic acid derivatives from the flower buds of Lonicera bournei. Planta Medica, 67(4), 322-355. [ Links ]
Yang, L., Fu, S., Khan, M. A., Zeng, W., & Fu, J. (2013). Molecular cloning and development of RAPD-SCAR markers for Dimocarpus longan variety authentication. SpringerPlus, 2, 501. [ Links ]
Zhang, H., Song, J., & Shi, J. (2011). Screen anti-influenza virus serum marker of Lonicera japonica by proteomics technology. Zhongguo Zhong Yao Za Zhi, 36(8), 1071-1074. [ Links ]
Bhat, A. A., Haniffa, M. A., Divya, P. R., Gopalakrishnan, A., Milton, M. J., Kumar, R., & Paray, B. A. (2012). Molecular characterization of eight Indian Snake-head species (Pisces: Perciformes Channidae) using RAPD markers. Molecular Biology Reports, 39(4), 4267-4273. [ Links ]
Chen, W. C., Liou, S. S., Tzeng, T. F., Lee, S. L., & Liu, I. M. (2012). Wound repair and anti-inflammatory potential of Lonicera japonica in excision wound-induced rats. BMC Complementary and Alternative Medicine, 12, 226. [ Links ]
Cho, W. K., Kim, H., Choi, Y. J., Yim, N. H., Yang, H. J., & Ma, J. Y. (2012). Epimedium koreanum Nakai water extract exhibits antiviral activity against porcine epidermic diarrhea virus in vitro and in vivo. Evidence Based Complementary and Alternative Medicine, 2012, 985151. [ Links ]
Dnyaneshwar, W., Preeti, C., Kalpana, J., & Bhushan, P. (2006). Development and application of RAPD- SCAR marker for identification of Phyllanthus emblica LINN. Biological & Pharmaceutical Bulletin, 29(11), 2313-2316. [ Links ]
Dutta, S. R., Kar, P. K., Srivastava, A. K., Sinha, M. K., Shankar, J., & Ghosh, A. K. (2012). Identification of RAPD and SCAR markers associated with yield traits in the Indian tropical tasar silkworm Antheraea mylitta drury. Genetics and Molecular Biology, 35(4), 743-751. [ Links ]
Dutta, S., Biswas, S., Mukherjee, K., Chakrabarty, U., Mallik, A., & Mandal, N. (2014). Identification of RAPD-SCAR marker linked to white spot syndrome virus resistance in populations of giant black tiger shrimp, Penaeus monodon Fabricius. Journal of Fish Diseases, 37(5), 471-480. doi: 10.1111/jfd.12128. [ Links ]
Frankham, R. (2008). Genetic adaptation to captivity in species conservation programs. Molecular Ecology, 17(1), 325-333. [ Links ]
Fu, J. J. (2012). Short protocols in medical molecular biology. Beijing: China Medical Science Press. [ Links ]
Fu, J., Yang, L., Khan, M. A., & Mei, Z. (2013). Genetic characterization and authentication of Lonicera japonica Thunb. by using improved RAPD analysis. Molecular Biology Reports, 40(10), 5993-5999. [ Links ]
Jiang, S., Jianhua, X., & Li, X. (2009). A Study on the RAPD and SCAR Molecular Markers of Piper Species. Journal of Agriculture and Rural Development in the Tropics and Subtropics, 110(2), 127-135. [ Links ]
Kumar, S., Sharma, R., Kumar, V., Govind, K., Vyas, G., & Rathore, A. (2013).Combining molecular-marker and chemical analysis of Capparis decidua (Capparaceae) in the Thar Desert of Western Rajasthan (India). Revista de Biología Tropical, 61(1), 311-320. [ Links ]
Kumla, S., Doolgindachbaporn, S., Sudmoon, R., & Sattayasai, N. (2012). Genetic variation, population structure and identification of yellow catfish, Mystus nemurus (C&V) in Thailand using RAPD, ISSR and SCAR marker. Molecular Biology Reports, 39(5), 5201-5210. [ Links ]
Lee, D. H., Lee, S. K., Lee, S. Y., & Lee, J. K. (2013). Development of SCAR Markers for the Identification of Phytophthora katsurae Causing Chestnut Ink Disease in Korea. Mycobiology, 41(2), 86-93. [ Links ]
Li, S. F., Tang, S. J., & Cai, W. Q. (2010). RAPD- SCAR markers for genetically improved NEW GIFT Nile Tilapia (Oreochrmis niloticus niloticus L.) and their application in strain identification. Zoological Research, 31(2), 147-153. [ Links ]
Liu, W., Li, L., Khan, M. A., & Zhu, F. (2012). Popular molecular markers in bacteria. Molecular Genetics, Microbiology & Virology, 27(3), 103-107. [ Links ]
Mei, Z., Fu, S., Yu, H., Yang L., Duan, C., Liu, Y., Gong, S., & Fu, J. (2014). Genetic characterization and authentication of Dimocarpus longan Lour. using an improved RAPD technique. Genetics and Molecular Research, 13(1), 1447-1455. [ Links ]
Mei, Z., Yang, L., Zhang, T., Yu, H., Gan, L., Zhang, L., Yang, M., & Fu, J. (2014). Improved RAPD Analysis of Canarium album (Lour.) Raeusch from Sichuan Province along Yangtze River in China. Annual Research & Review in Biology, 4(1), 51-60. [ Links ]
Noormohammadi, Z., Hasheminejad-Ahangarani, F. Y., Sheidai, M., Ghasemzadeh-Baraki, A. S., & Alishah, O. (2013). Genetic diversity analysis in Opal cotton hybrids based on SSR, ISSR, and RAPD markers. Genetics and Molecular Research, 12(1), 256-269. [ Links ]
Park, H. S., Park, K. I., Lee, D. H., Kang, S. R., Nagappan, A., Kim, J. A., Kim, E. H., Lee, W. S., Shin, S. C., Hah, Y. S., & Kim, G. S. (2012). Polyphenolic extract isolated from Korean Lonicera japonica Thunb. Induce G2/M cell cycle arrest and apoptosis in HepG2 cells: involvements of PI3 K/Akt and MAPKs. Food and Chemical Toxicology, 50(7), 2407-2416. [ Links ]
Rajesh, M. K., Jerard, B. A., Preethi, P., Thomas, R. J., Fayas, T. P., Rachana, K. E., & Karun, A. (2013). Development of a RAPD-derived SCAR marker associated with tall-type palm trait in coconut. Scientia Horticulturae, 150, 312-316. [ Links ]
Shakeel, M., Ilyas, M., & Kazi, M. (2013). Evaluation of synthetic hexaploid wheats (derivative of durum wheats and Aegilops tauschii accessions) for studying genetic diversity using randomly amplified polymorphic DNA (RAPD) markers. Molecular Biology Reports, 40(1), 21-26. [ Links ]
Williams, J. G., Kubelik, A. R., Livak, K. J., Rafalski, J. A., & Tingey, S. V. (1990). DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Research, 18(22), 6531-6535. [ Links ]
Xiang, T., Xiong, Q. B., Ketut, A. I., Tezuka, Y., Nagaoka, T., Wu, L. J., & Kadota, S. (2001). Studies on the hepatocyte protective activity and the structure- activity relationships of quinic acid and caffeic acid derivatives from the flower buds of Lonicera bournei. Planta Medica, 67(4), 322-355. [ Links ]
Yang, L., Fu, S., Khan, M. A., Zeng, W., & Fu, J. (2013). Molecular cloning and development of RAPD-SCAR markers for Dimocarpus longan variety authentication. SpringerPlus, 2, 501. [ Links ]
Zhang, H., Song, J., & Shi, J. (2011). Screen anti-influenza virus serum marker of Lonicera japonica by proteomics technology. Zhongguo Zhong Yao Za Zhi, 36(8), 1071-1074. [ Links ]
1. Research Center for Preclinical Medicine, Luzhou Medical College, Luzhou, Sichuan Province 646000, China; 690199186@qq.com, asadkhanbmj@yahoo.com, xuguangyin1@163.com, yangman0205@126.com, 1517440878@qq.com, weichunli2013@163.com, 47191096@qq.com, 871345204@qq.com, longyan0124@foxmail.com
2. Research Center for Preclinical Medicine, Luzhou Medical College, Luzhou, Sichuan Province 646000 China, and Forensic Center, Luzhou Medical College, Luzhou, Sichuan 646000, China; fujunjiang@hotmail.com
2. Research Center for Preclinical Medicine, Luzhou Medical College, Luzhou, Sichuan Province 646000 China, and Forensic Center, Luzhou Medical College, Luzhou, Sichuan 646000, China; fujunjiang@hotmail.com
Received 17-II-2014. Corrected 22-V-2014. Accepted 25-VI-2014.