Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO -
 Access statistics
Access statistics
Related links
-
 Similars in
SciELO
Similars in
SciELO  uBio
uBio
Share
Revista de Biología Tropical
On-line version ISSN 0034-7744Print version ISSN 0034-7744
Rev. biol. trop vol.55 n.2 San José Jun. 2007
Molecular phylogenetic relationships among four species of the mangrove tree genus Bruguiera (Rhizophoraceae), as revealed by chromosome and RAPD markers
Pragnya Sahoo, Satyanarayan Jena, Suprava Mohanty & Anath Bandhu Das*
Cytogenetics Laboratory, Regional Plant Resource Centre, Nayapalli, Bhubaneswar-751015, Orissa, India, Fax: ++-91-674-2550274; * a_b_das@hotmail.com
Received 02-VII-2003. Corrected 09-I-2006. Accepted 13-X-2006.
Abstract: Analysis of karyotype, nuclear DNA content and RAPD markers were performed in four species of Bruguiera (Rhizophoraceae) of Bhitarkanika mangrove forests, Orissa, India. Detailed karyotype analysis revealing 2n=34 in B. cylindrica and 2n=36 in B. gymnorrhiza was reported for the first time and 2n=34 in B. parviflora and B. sexangula was confirmed. On the basis of the common types of chromosomes present among Bruguiera, two distinct groups were found; one consists of B. cylindrica and B. parviflora and the other of B. gymnorrhiza and B. sexangula. The symmetrical karyotype with same chromosome types grouped B. cylindrica and B. parviflora together and presence of Type E chromosomes placed B. gymnorrhiza and B. sexangula in a separate group, suggesting their closer affinity in their respective group. Analysis of chromosome length, volume, INV and 4C DNA content confirmed this division. Nuclear DNA content was two-fold higher (~17.0 pg) in the second group than in the first (~8.0 pg). The amplification products generated through RAPD revealed 1-9 amplicons with size variations from 600 bp to 2 500 bp with 49.31% genetic similarity between B. gymnorrhiza and B. sexangula and 47.10% in between B. cylindrica and B. parviflora. The high copy number marker band (~ 1 100 bp) yielded in OPN-15 primer in B. parviflora the characteristic DNA marker, which was cloned and used as probes for assessment of genetic diversity, and demonstrated its close genetic affinity to B. cylindrica. B. gymnorrhiza and B. sexangula also produced similar marker bands of ~600 bp and ~2 200 bp in the same primer. All of the cytological, 4C DNA content and RAPD data confirmed the existence of two taxonomically distinct groups of Bruguiera: one consisting of B. cylindrica and B. parviflora and the other of B. gymnorrhiza and B. sexangula as placed earlier (1862) in the tribe Rhizophoreae by Bentham and Hooker, on the basis of the flowering habits of Bruguiera. Genetically, the B. sexangula and B. gymnorrhiza group was found to be very closely, rather than distantly, related to B. parviflora and B. cylindrica. Our results demonstrate that molecular markers together with cytological evidence provide an effective tool to access the existing interspecific genetic polymorphism in mangrove species, to solve the taxonomic problems and to design their conservation strategy. Rev. Biol. Trop. 55 (2): 437-448. Epub 2007 June, 29.
Key words: 4C DNA content, Bruguiera, molecular phylogeny, tree mangrove, karyotype.
Mangroves are perennial plants found in the intertidal zones of coastal wetlands in tropical regions; many species that are found exclusively there are called true mangroves and are capable of forming dense pure stands of trees. The mangrove ecosystem plays an important role in maintaining coastal ecological balance and is also one of the most highly productive and dynamic ecosystems to mankind, providing food, livelihood and eco logical security.
The genus Bruguiera, a true mangrove, of the family Rhizophoraceae consists of four spe cies distributed throughout the coastal regions of India: B. cylindrica, B. gymnorrhiza, B. parviflora and B. sexangula. The fruits of B. parviflora are used in making medicines for eye diseases, the bark of B. gymnorrhiza is used in the treatment of diarrhea and the leaves of B. cylindrica are used as fodder (Naskar and Mandal 1999). These mangroves are now endangered because they are being exploited indiscriminately for their timber (Das et al. 2002) which is resistant to post harvest pest infection due to the high deposits of tannin in the wood. Ignorance and negligence of its importance has led to a drastic reduction worldwide and the local extinction of many species and populations of Bruguiera.
Cytological and molecular phylogenetic research has mainly focused on the analysis of genomes of different taxa, but the detailed studies on the cytological, cytochemical and molecular aspects of Bruguiera are lacking. However, chromosome and cytophotometric studies are extremely difficult in these species due to the accumulation of a high amount of secondary metabolites and their tiny chromosomes. Studies of genetic variability and interspecific relationships among the mangrove species using molecular markers have been used to accurately quantify the extent of genetic diversity among the species (Waugh and Powell 1992). Unlike morphological markers, molecular markers are not prone to environmental influences and accurately portray the genetic relationships between plant groups (Emmarold et al. 2001). These markers can also be used to select priority areas for conservation and provide vital information to improve strategies for genetic conservation (New-bury and Ford-Lloyd 1993). The somatic chromosome number 2n=34 in B. parviflora and B. sexangula was reported earlier by our group (Das et al. 1995) and is in contrast with an earlier report of 2n=36 in B. parviflora (Sidhu 1968). Scanty reports on RAPD markers on Bruguiera were noted (Parani et al. 1998) without any report on B. parviflora, B. gymnorrhiza and B. sexangula.
Bentham and Hooker (1862) placed Bruguiera in the tribe Rhizophoreae along with the genera Rhizophora, Ceriops and Kandelia. The genus Bruguiera was found to be distinct in its explosive mechanism of pollen release, which is largely triggered by visiting pollinators, a phenomenon not observed in other genera of this tribe. The genus was further segregated into two distinct taxonomic groups of species based on flowering differences: the large, solitary-flowering group consisting of B. gymnorrhiza and B. sexangula and the small multi-flowering group of B. parviflora and B. cylindrica. The difference in flower size also correlated with differences in leaf and fruit size. However, Ding Hou (1958) pointed out that the distinction between the two groups was obscured by the later discovery of the two species, B. hainesi and B. exarillata, which have some morphologically intermediate characteristics. The first species sometimes has solitary flowers, the second sometimes has flowers in pairs and the size difference between flowers is slight; however, there is a fairly sharp distinction between the species on the basis of flower orientation in relation to different types of pollinators (birds vs. insects). Despite this variation, there is not sufficient justification to revise the classification.
The present study deals with detailed karyotype analysis, estimation of 4C DNA content and RAPD polymorphism analysis in the four species of Bruguiera belonging to two morphological groups of the tribe Rhizophoreae in order to ascertain their genetic diversity and to establish phylogenetic relationships as well as affinities, if any, of the four species of Bruguiera belonging to the two taxonomic groups and to resolve the taxonomic status of these genera using the existing taxonomic classification.
Materials and methods
Propagules of Bruguiera parviflora Roxb., B. cylindrica (Linn.) Bl., B. gymnorrhiza (Linn.) Lamark and B. sexangula (Lour.) Poir. were collected from Bhitrarkanika, Orissa and were grown in the experimental mangrove nursery at Regional Plant Resource Centre, Bhubaneswar and voucher specimens of all the species were kept in the herbaria of the centre.
Karyotype analysis: for the somatic chromosome study, healthy fresh root-tips were pretreated in ¼th saturated solution of paradichlorobenzene (pDB) and aesculine for 2½ h at 14oC followed by overnight fixation in lactopropionic alcohol (1:3). The root-tips were stained in 2% lactopropionic orcein after cold hydrolysis in 5N HCl for 10 min and were squashed in 45% lactopropionic acid. Five well scattered metaphase plates were selected for karyotype analysis of each species. The total chromosome length was ascertained by adding the length of all chromosomes in the karyotype and the total chromosome volume of a karyotype was calculated by applying the formula πr2h, where r and h represent the radius and length of the chromosome, respectively. The total form percentage (TF%) of a karyotype was the average of the sum total F% of a karyotype. The mean values of total genomic chromosome length and total chromosome volume with standard error were calculated.
Nuclear DNA content: for Feulgen cytophotometric estimation of 4C DNA content, ten fixed root-tips from each species were hydrolyzed in 1N HCl for 12 min at 60oC, washed in distilled water and rinsed in SO2 water (Fox 1969). Root tips were stained in Schiffs reagent for 2h at 14oC and each root-tip squash was prepared in 45% lactopropionic acid. In situ nuclear DNA content was estimated from metaphase chromosomes using a Nikon Optiphot microscope fitted with a microspectrophotometer using monochromatic light at 550 nm following the method of Sharma and Sharma (1980), with ten scorings made from each slide. In situ DNA content was obtained on the basis of optical density, which was converted to picograms (pg) using the 4C nuclear DNA values (67.1 pg) for Allium cepa var. Deshi (Vant Hof 1965) as a standard, whose root tips were prepared and scored in the same way as the experimental material. The correlation coefficient analysis was done between different chromosomal parameters to compare genomic characteristics. Analysis of variance (ANOVA) was performed among the nuclear DNA values using Duncans multiple range test (Harter 1960).
Interphase nuclear volume (INV): for the scoring of interphase nuclear volume (INV), root-tips about 2-2.5 mm in length were fixed in 1:3 acetic acid: ethanol for 24 h at 25oC, hydrolyzed in 1N HCl at 4oC for 15 min. After a thorough washing, the root-tips were put into Schiffs reagent for 1 h at 20oC and kept in the dark for staining. Squash preparation was done in 45% lactopropionic acid. Scoring was done following the method of Das et al. (1993).
Isolation of nuclear DNA: Genomic DNA was isolated from young expanding leaves (Saghai-Maroof et al. 1984). Young leaves (10 g) were ground to fine powder in liquid nitrogen and suspended in three volumes of suspension buffer (pH 8) containing 50 mM EDTA, 100 mM Tris-HCl, 0.8 M NaCl, 0.5 M sucrose, 2% Triton-X 100, 0.1% β-mercaptoethanol and incubated at 60 oC for 30 min. The suspension was centrifuged at 10 000 g for 10 min at room temperature. Then the pellet was extracted in 20 ml of extraction buffer containing 2 mM EDTA, 100 mM Tris-HCl, 1.5M NaCl, 2% CTAB, 1% β-mercaptoethanol and incubated at 60oC for 30 min and centrifuged again at 10 000 g for 20 min at room temperature. The aqueous phase was transferred to a new 50 ml tube and the DNA was precipitated with a double volume of chilled isopropanol, and hooked out and dried with vacuum drier and dissolved in a minimum amount of TE (10 mM Tris-HCl, 1 mM EDTA; pH 8). The DNA was again purified by treating with RNase at 37oC for 1 h followed by chloroform: isoamyl alcohol extraction and ethanol precipitation in the presence of 0.3M sodium acetate (pH 5.2). The DNA was spooled out, washed in 70% ethanol, air-dried and dissolved in T10E1 buffer and DNA concentration was estimated using Versafluor TM Fluorometer (Bio-Rad, USA) using Hoechst 33258 as the fluorimetric dye. The DNA was diluted to a final concentration of 25 ng µl-1 using T10E1 buffer as template for RAPD analysis.
RAPD analysis: RAPD profiles were generated using single decamer primers (Operon Technologies, Alameda, USA) in polymerase chain reaction (PCR) following the standard protocol of Williams et al. (1990). Each reaction mixture (25 µl) for PCR amplification, was prepared with 25 ng genomic template DNA, 200 µM each of dNTP, 25 ng primer, 0.5 unit Taq DNA Polymerase (Bangalore Genei Pvt. Ltd., Bangalore, India) and 10x PCR assay buffer (50 mM KCl, 10 mM Tris-HCl, 1.5 mM MgCl2, pH 9.0). The PCR reaction was carried out in a GeneAmp PCR 2400 thermal cycler (Perkin Elmer, USA) programmed for 45 cycles using 10 primers each for all the species. The first cycle consisted of denaturation at 94oC for 5 min, primer annealing at 37oC for 1min and DNA polymerization at 72oC for 2 min. In the next 44 cycles the period of denaturation was maintained at 1min while the primer annealing and DNA polymerization were kept the same as in the first cycle. The last cycle consisted of only primer extension (72oC) for 8 min. The amplified samples were stored at 4oC and separated by electrophoresis in 1.5% agarose gel in 1x TAE buffer for 3 h at 55 V. Gene Ruler 100 bp DNA ladder plus (MBI Fermantas, Lithuania) was used as a marker to determine the size of the amplicons. Amplified products were visualized by staining the gel with ethidium bromide, were photographed, and an image was captured using an image analyzer Gel Doc-G 700 (Bio-Rad, USA) for documentation and data analysis. Only those amplification products that appeared consistently in triplicate were scored for further analysis.
Statistical analysis
Chromosome and nuclear DNA analysis: mean values of total genomic chromosome length and volume with standard error were calculated. The ANOVAs were performed on the nuclear DNA content using Duncans multiple range test (Harter 1960).
RAPD data analysis: in RAPD analysis, the presence or absence of the bands were taken into consideration and the difference in the intensity of the bands was ignored. From RAPD data a binary matrix was obtained and calculate using the multivariate analysis program NTSYS-PC (Rohlf 1993). The binary matrix was transformed in a similarity matrix using the Jaccards coefficient. From this matrix a phylogenetic dendrogram was obtained by cluster analysis following the unweighted pair group with arithmetic mean (UPGMA) method (Sneath and Sokal 1973), using NTSYS version 1.7, Exeter Software, New York, USA.
Results
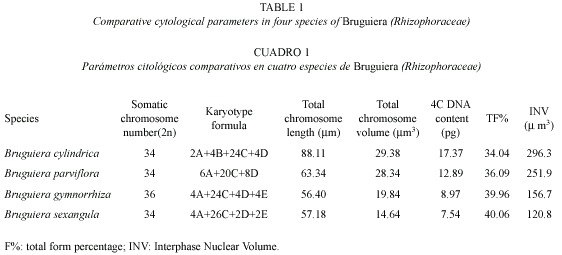
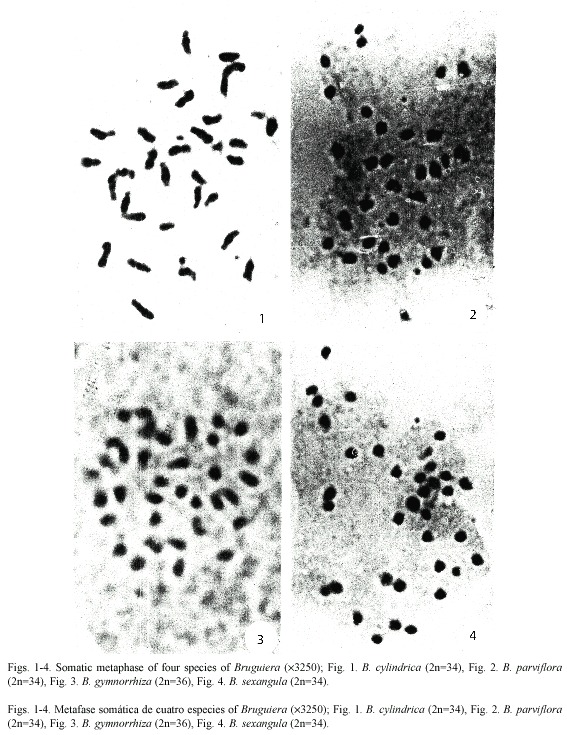
Karyotype analysis: The somatic chromosome number varied from 2 n = 34 in B. cylindrica, B. parviflora and B. sexangula to 2 n = 36 in B. gymnorrhiza. Chromosome structure was highly similar among species types with minor structural variations in the karyotypes. On the basis of size and position of the primary and secondary constrictions, five types of chromosomes A, B, C, D and E were identified in the four species, although the species differed from one another in chromosome types and/or copy number. Type A chromosomes are medium-sized with primary and secondary constrictions at the nearly median and nearly sub-terminal positions. Type B chromosomes have two constrictions, one median and the other sub-terminal, producing a satellite body on the long arm. Types C and D chromosomes are medium to small in size with median and sub-median constrictions respectively. Type E chromosomes are very small in size with a primary constriction close to the submedian position. A detailed description of the somatic complements and different genomic characteristics of the four species studied of Bruguiera showed species specific variations in the genomic behavior (Table 1, Figs. 1-4). Type A, C and D chromosome were present in all the species of Bruguiera in different doses and were the only chromosome types present in B. parviflora; only type B was obtained in B. cylindrica and Type E chromosomes were observed in B. gymnorrhiza and B. sexangula. The total chromosome length varied from 56.40 µm in B. gymnorrhiza to 88.11 µm in B. cylindrica. Minimum chromosome volume was 14.64 µm3 in B. sexangula and maximum 29.38 µm3 in B. cylindrica. The total percentage (TF%) varied from 34.04% in B. cylindrica to 40.06% in B. sexangula (Table 1).
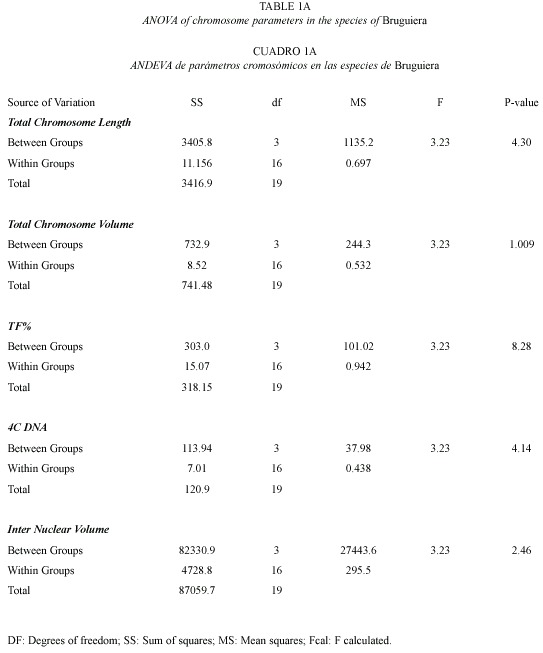
4C Nuclear DNA and INV: The 4C nuclear DNA amount varied significantly from 7.45 pg in B. sexangula to 17.37 pg in B. cylindrica. The average DNA amount per chromosome also varied from species to species, i.e. 0.21 pg in B. sexangula, 0.26 pg in B. gymnorrhiza, 0.37 pg in B. parviflora, 0.51 pg in B. cylindrica. The frequency polygraph of the INV showed a distinct peak around the mean value in all the species (data not shown). Correlation coefficient analysis showed a significant correlation of total chromosome length with volume (0.732), 4C DNA content (0.939) and INV (0.850). 4C DNA content showed positive correlation to chromosome volume (0.921) and INV (0.979). The ANOVA and Duncans multiple range tests showed that the variation in the nuclear DNA amounts among the different species of Bruguiera were significant at the 1 % level (Table 1a).
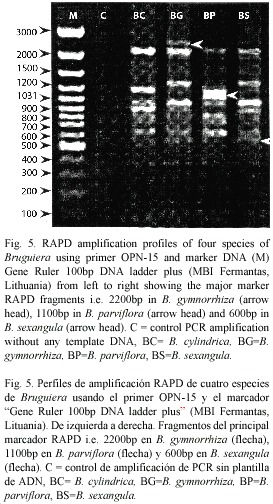
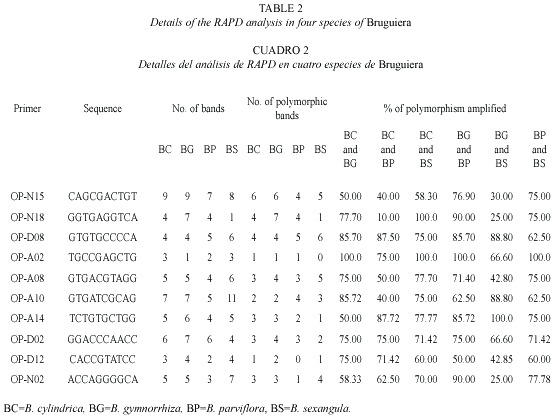
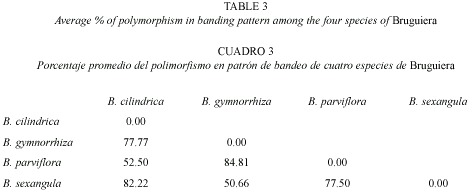
RAPD analysis: The RAPD profiles were reproducible and displayed no variation within the different plants studied in triplicate. On the basis of RAPD analysis results, interspecific variations were reflected in their genomic DNA amplified with ten primers. The amplification products in RAPD ranged from one to nine in the four species, with fragment size ranging from 0.60 KB to 2.5 KB (Fig. 5). The total number of amplification products in all the species in each primer ranged from nine in OPA-2 to 33 in OPN-15 (Table 2). The maximum at 11 amplified products were obtained in B. sexangula with OPA-10 primer. Different primers responded differently according to the genomic characteristics of the species and the numbers of amplicon varied from 7-11 bands in OPA-10, 7-9 bands in OPN-15 and 4-7 bands in OPD-2 primer. The maximum of 36 polymorphic bands was obtained in B. gymnorrhiza, followed by 30 in B. cylindrica, 28 in B. sexangula and 27 in B. parviflora. A maximum of six polymorphic bands could be resolved in B. cylindrica, B. gymnorrhiza and B. sexangula using OPN-15 and OPD-8 primers whereas five polymorphic bands were obtained in B. parviflora in OPD-8 primer. The maximum average percentage of polymorphism (84.81%) was observed between B. parviflora and B. gymnorrhiza, whereas the minimum DNA polymorphism (50.66%) existed between B. gymnorrhiza and B. sexangula (Table 3).The RAPD banding pattern obtained from the primers OPN-18 and OPA-2 revealed 100% polymorphism between B. cylindrica and B. sexangula as well as between B. gymnorrhiza and B. parviflora.
Cluster analysis: The cluster analysis of the RAPD profiles of all the primers follow ing the method of unweighted pair group with arithmetic mean (UPGMA) revealed four species of Bruguiera distributed into two distinct branches of a single tree (Fig. 6); B. cylindrica and B. parviflora were clustered together with a similarity coefficient of 0.45 and B. gymnorrhiza and B. sexangula with similarity coefficient 0.83.
Discussion
Karyotype, genome length and INV: Detailed karyotype analysis of four species of Bruguiera revealed 2 n = 34 diploid chromosomes in all the species except B. gymnorrhiza (2 n = 36). Although the chromosome numbers were same in B. cylindrica, B. parviflora and B. sexangula, they differed in the type and number of secondary constricted chromosomes in their karyotype. The chromosome number reports 2n=34 in B. cylindrica and B. parviflora reconfirm our earlier report (Das et al. 1995) and the diploid number in B. gymnorrhiza (2 n = 36) and in B. sexangula (2 n = 34) was reported for the first time. Among the species studied, the three chromosome types A, C, D were present in all four species. Type B chromosome was only characteristic of B. cylindrica with secondary constriction on the long arm of the chromosomes and Type E chromosomes were present in B. gymnorrhiza and B. sexangula in different doses. B. cylindrica and B. parviflora had six secondary constricted chromosomes, with chromosome types A and B present in B. cylindrica, and a high number of D type chromosomes in B. parviflora. Four Type A chromosomes were present in both B. gymnorrhiza and B. sexangula. The median constricted chromosomes were nearly equal in all the species studied. Evidently, structural changes as well as changes in the amount of heterochromatin might have played a vital role in differentiating at species level (Das et al. 1995).
The TF % values varied from 34.96% to 40.06% among the four species might be due to structural alterations in the genome during speciation revealed by a duplication of the parts of some chromosomes, or by translocations, some with secondary constrictions and some without (Das et al. 2001). TF% values of about 35% in B. cylindrica and B. parviflora and ~40% in B. gymnorrhiza and B. sexangula suggest two distinct groups; the former had more sub-median chromosomes while the latter presented more median chromosomes. The gradual decrease of sub-metacentric chromosomes in the karyotypes of first group (B. cylindrica and B. parviflora) and an increase in the number of median chromosomes in the second group (B. sexangula and B. gymnorrhiza) suggests a gradual shifting of primary chromosome constriction through translocation or alteration and rearrangements of chromosomal arms in the evolution. Furthermore, the symmetrical karyotype with same number of secondary constricted chromosomes in B. cylindrica, B. parviflora, B. gymnorrhiza and B. sexangula suggests their close affinity.
The total chromosome length and total chromosome volume varied significantly e.g., in B. gymnorrhiza with 56.40 µm and 19.84 µm3, respectively and in B. cylindrica with a length of 88.11 µm and a volume of 29.38 µm3. The first time reports of 4C DNA in these species of Bruguiera classified each in one of the two established groups; one group consisting of B. gymnorrhiza and B. sexangula had ~7 pg to 8 pg of DNA and the other consisting of B. cylindrica and B. parviflora had about twice the DNA content. The average DNA content was highly correlated to average chromosome volume and INV. However, in the eukaryotic system, chromosome volume is not only determined by its DNA, but also by its basic and non basic proteins as well. The highest DNA content found was 17.37 pg in B. cylindrical, which had all the types of chromosomes except Type E and the lowest was found in B. sexangula with 7.54 pg, which had two Type E chromosomes but lacked any Type B. The variability in DNA amount has often been attributed to loss or addition of highly repetitive DNA sequences rather than to the AT-or GC-rich sequences in the genome (Martel et al. 1997). These highly repetitive DNA sequences reach a certain level and become stabilized during microevolution and gradual selection (Price et al. 1980). Such interspecific variation in DNA amount is not unique to angiosperm species (Laurie and Bennet 1985, Rayburn et al. 1989). B. cylindrical, along with B. parviflora, has adapted to the comparatively low saline areas in the mangrove community, while B. sexangula and B. gymnorrhiza have adapted to the comparatively high saline zones of mangroves, suggesting species adaptability through chromosomal structural alterations and their resulting changes at the DNA level during the course of evolution.
RAPD profile: The previous studies on the genetic structure of the four Bruguiera species was restricted largely to chromosome number and karyotype (Das et al. 1995). Parani et al. (1998) reported the RAPD analysis on only the B. cylindrica species, but no such studies had been done on the other three species B. parviflora, B. gymnorrhiza and B. sexangula. In the present study, the RAPD analysis of the four species of Bruguiera that form mangrove forests in Orissa, recorded the highest average percentage of polymorphism (84.81%) between B. gymnorrhiza and B. parviflora and the lowest between B. gymnorrhiza and B. sexangula (50.66%). This suggests a greater similarity in genetic constitution between B. gymnorrhiza and B. sexangula as compared to B. parviflora and B. cylindrica. The maximum percentage of similarity (49.31%) in banding pattern was observed between B. gymnorrhiza and B. sexangula, followed by B. cylindrica and B. parviflora (47.1%). The UPGMA cluster analysis of the four species showed a high coefficient of interspecific similarity among the species, forming the two distinctly separate groups, B. cylindrica and B. parviflora (0.45) and B. gymnorrhiza and B. sexangula (0.57), also confirmed by the karyotype, 4C DNA content and Interphase Nuclear Volume analysis.
Mangrove species are constantly subject to physiological stress caused by fluctuating growing conditions (Chapecker 1994). Despite such extremes, they have sucessessfully colonized suitable areas through morphological, physiological and reproductive adaptations (Clough 1994). Therefore, depending on their edaphic preferences and adaptation, different species are likely to display varying degrees of polymorphism. The observations on the RAPD analysis of Bruguiera are in accordance with the cytological studies with a genomic similarity of 49.31% between B. gymnorrhiza & B. sexangula and of 47.10% between B. cylindrica and B. parviflora. The OPA-2 primer yielded maximum polymorphism in all the species at a level of 100% between B. cylindrical & B. sexangula and B. gymnorrhiza and
B. parviflora. The primer OPN-15 showed a significant marker band at 1 100 bp with a high copy number in B. parviflora, which was the most prominent marker band. B. gymnorrhiza produced an unique marker band at 2 200 bp DNA which differentiates this species from the others. B. gymnorrhiza and B. sexangula also produced similar marker bands at 600 bp. The observed interspecific differences could be ascribed to the fluctuating climatic habitat conditions and may be species specific. The distinct RAPD bands could be cloned and used as markers for species identification. In conclusion, our results demonstrate that molecular markers provide an effective tool to access the existing interspecific genetic polymorphism in mangrove species and to design their conservation strategy. The two taxonomically and genetically distinct groups of Bruguiera consist of B. cylindrica and B. parviflora in one group and B. gymnorrhiza and B. sexangula in the other. All the karyotypic, DNA content and RAPD marker analysis suggests and confirms the taxonomic segregation of these groups in the Rhizophoreae tribe as suggested by Bentham and Hooker (1862) on the basis of the flowering habits of Bruguiera. Genetically, the group consisting of B. sexangula and B. gymnorrhiza was more closely, than distantly, related to B. parviflora and B. cylindrica. A better understanding of these groups could be of phylogenetic interest.
Acknowledgment
The authors are thankful to the Ministry of Forest and Environment, Government of India Grant No. 3/7/2000-C3 (M) for financial assistance.
Resumen
Estudiamos cuatro especies del mangle Bruguiera (Rhizophoraceae) en Orissa, India. Los cromosomas indican queB. cylindrica y B. parviflora son un grupo taxonómico, y que B. gymnorrhiza y B. sexangula son otro. Genéticamente, el par B. sexangula y B. gymnorrhiza está cercanamente emparentado con B. parviflora and B. cylindrica. Nuestros datos indican que el uso combinado de marcadores genéticos y evidencia citológica permiten discernir el polimorfismo genético interespecífico en los mangles, tanto para resolver problemas taxonómicos como para diseña estrategias eficaces de conservación.
Palabras clave: contenido de ADN4C, Bruguiera, filogenia molecular, mangle, cariotipo.
References
Bentham, G. & J.D. Hooker. 1962. Rhizophoreae: Genera Plantarum Vol. 1. Linnean Society, London. 677 p. [ Links ]
Clough, B.S. 1994. Climatic impacts on mangrove ecosystems. p. 39-43. In V. Deshmukh & V. Balaji (eds.). Conservation of mangrove genetic resources: Training manual. ITTO-CRSARD Project, MSSRF, Madras, India. [ Links ]
Das, A.B. & R. Mallick. 1993. Karyotype diversity and interspecific 4C DNA variation in Bupleurum. Biologia Plantarum 35: 355-363. [ Links ]
Das, A.B., U.C. Basak & P. Das. 1995. Chromosome number and karyotype diversity in the Rhizophoraceae found in the mangrove forest of Orissa. Cytobios 81: 27-35. [ Links ]
Das, A.B., A.K. Mukhejee & P. Das. 2001. Molecular phylogeny of Heritiera Aiton (Sterculiaceae), a tree mangrove, variations in RAPD markers and nuclear DNA content. Bot. J. Linn. Soc. 136: 221-229. [ Links ]
Emmarold, E.M., H.M. Sinclair & B. Mark. 2001. Use of random amplified polymorphic DNA (RAPD) markers to reveal genetic diversity within and between populations of cashew (Anacardium occidentale L.). J. Hortic. Sci. Biotech. 76: 375-383. [ Links ]
Fox, D.P. 1969. Some characteristics of the cold hydrolysis technique for staining plant tissues by the Feulgen reaction. J. Histochem. Cytochem. 17: 266-272. [ Links ]
Harter, H.L. 1960. Critical values for Duncans Multiple range test. Biometrics 16: 671-685. [ Links ]
Laurie, D.A. & M.D. Bennet. 1985. Nuclear DNA content in the genera Zea and Sorghum. Intergeneric, interspecific and intraspecific variation. Heredity 55: 307-313. [ Links ]
Martel, E., D. Denay, S. Siljakyakovlev, S. Brown & A. Sarr. 1997. Genome size variation and basic chromosome number in pearl millet and fourteen related Pennisetum Species. J. Heredity 88: 139-143. [ Links ]
Naskar, K. & R. Mandal. 1999. Ecology and Biodiversity of Indian Mangroves, (Vol-II), Daya, Delhi, India. 364 p. [ Links ]
Parani, M., M. Lakshmi, P. Senthilkumar, R. Nivedita & A. Parida. 1998. Molecular phylogeny of mangroves V. Analysis of genome relationships in mangrove species using RAPD and RFLP markers. Theor. App. Gen. 97: 617-625. [ Links ]
Price, H. J., K. Bachaman, K.L. Chambers & J. Riggs. 1980. Detection of interspecific variation in nuclear DNA content in Microseris douglasii. Bot. Gaz. 141: 195-198. [ Links ]
Rayburn, A.L., J.A. Auger, E.A. Benzinger & A.G. Hepburn. 1989. Detection of interspecific DNA content variation in Zea mays L. by flow cytometry. J. Exp. Bot. 40: 1179-1183. [ Links ]
Rohlf, F.J. 1993. NTSYS-pc. Numerical and reproductive adaptations of Australian mangroves. p. 232-238. In Chapman V. J. (ed.) Ecosystems of the world. Vol. 1: Wet Coastal Ecosystems. Elsevier. Amsterdam, Netherlands. [ Links ]
Saghai–Maroof, M.A., K.M. Soliman, H.A. Jorgensen & H.A. Allard. 1984. Ribosomal DNA spacer length polymorphism in Barley. Mendelian inheritance, chromosomal locations and population dynamics. Proc. Nat. Acad. Sci. USA 81: 8014-8018. [ Links ]
Sharma, A.K. & A. Sharma. 1980. Chromosome techniques: theory and practice. Butterworths. London, United Kingdom. 575p. [ Links ]
Sidhu, S.S. 1968. Further studies on the cytology of mangrove species in India. Caryologia 21: 353-357. [ Links ]
Sneath, P.H.A. & R. Sokal. 1973. Numerical taxonomy. Freeman. San Francisco, California, USA. 359p. [ Links ]
Vant Hof, J. 1965. Relationships between mitotic cycle duration and the average rate of DNA synthesis in the root meristem cells of sevaral plants. Exp. Cell Res. 39:48. [ Links ]
Waugh, R. & W. Powell. 1992. Using RAPD markers for crop improvement. Trends Biotech. 10:186-191. [ Links ]
Williams, J.G.K., A.R. Kubelik, K.J. Livak, J.A. Rafalski & S.V. Tingey. 1990. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic Acid Res. 18:6531-6535. [ Links ]