Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO -
 Access statistics
Access statistics
Related links
-
 Similars in
SciELO
Similars in
SciELO  uBio
uBio
Share
Revista de Biología Tropical
On-line version ISSN 0034-7744Print version ISSN 0034-7744
Rev. biol. trop vol.52 n.3 San José Sep. 2004
Diversity of Bacillus thuringiensis strains isolated from coffee plantations infested with the coffee berry borer Hypothenemus hampei
Glen Arrieta1 , Alejandro Hernández1 & Ana M. Espinoza1,2
1 Centro de Investigación en Biología Celular y Molecular (CIBCM). Ciudad de la Investigación, Universidad de Costa Rica. San José, Costa Rica
2 Escuela de Agronomía, Facultad de Ciencias Agroalimentarias, Universidad de Costa Rica, San José. Costa Rica.
Received 10-III-2004. Corrected 18-VIII-2004. Accepted 25-VIII-2004.
Abstract
The coffee berry borer Hypothenemus hampei Ferrari (Coleoptera: Scolytidae) was first reported infecting Costa Rican coffee plantations in the year 2000. Due to the impact that this plague has in the economy of the country, we were interested in seeking new alternatives for the biological control of H. hampei, based on the entomopathogenic bacteria Bacillus thuringiensis. Atotal of 202 B. thuringiensis isolates obtained from Costa Rican coffee plantations infested with H. hampei were analyzed through crystal morphology of the crystal inclusions and SDS-PAGE of d-endotoxins, while 105 strains were further evaluated by PCR for the presence cry, cyt and vip genes. Most of the Bt strains showed diverse crystal morphologies: pleomorphic (35%), oval (37%), bipyramidal (3%), bipyramidal and oval (12%), bipyramidal, oval and pleomorphic (10%) and bipyramidal, oval and cubic (3%). The SDS-PAGE analyses of the crystal preparations showed five strains with d -endotoxin from 20 to 40 kDa, six from 40 to 50 kDa, seven from 50 to 60 kDa, 19 from 60 to 70 kDa, 29 from 70 to 100 kDa and 39 from 100-145 kDa. PCR analyses demonstrated that the collection showed diverse cry genes profiles having several genes per strain: 78 strains contained the vip3 gene, 82 the cry2 gene, 45 the cry1 and 29 strains harbored cry3-cry7 genes. A total of 13 strains did not amplified with any of the cry primers used: cry1, cry2, cry37, cry5, cry11, cry12 and cry14. Forty-three different genetic profiles were found, mainly due to the combination of cry1A genes with other cry and vip genes. The genetic characterization of the collection provides opportunities for the selection of strains to be tested in bioassays against H. hampei and other insect pests of agricultural importance. Rev. Biol. Trop. 52(3): 757-764. Epub 2004 Dic 15.
Key words: Bacillus thuringiensis, crystals, cry genes, d -endotoxins, Hypothenemus hampei, coffee.
Palabras clave: Bacillus thuringiensis, cristales, genes cry, d -endotoxinas, Hypothenemus hampei, café.
Coffee is an important cash crop for several Latin American countries, including Costa Rica. Coffee production has several phytosanitary constrains however, the most important pest is the coffee berry borer Hypothenemus hampei Ferrari (Coleoptera: Scolytidae). This insect attacks coffee beans in all maturation stages, where it carries out most of its life cycle, causing important yield and grain quality loses (Le Pelley 1968). The most common strategy for the control of this insect is based on cultural practices and the application of synthetic insecticides such as endosulphan, known for being harmful to the environment, contaminating superficial and underground water, as well as for its toxicity to animals, beneficial insects and human health. In addition, it has been demonstrated that the indiscriminate use of this insecticide induces the emergence of resistant insect populations of H. hampei (Brun et al. 1989). As a consequence the search for new alternatives for the sustainable control of H. hampei is of great environmental impact.
Several entomopathogenic microorganisms are used for the biological control of insect pests and one of the most widely used is Bacillus thuringiensis (Bt) (Samsonov et al. 1997). Bacillus thuringiensis is a Gram-positive bacterium that synthesizes during sporulation crystal inclusions constituted by d -endotoxins of insecticidal activity, encoded by the cry and cyt genes. These d -endotoxins have molecular weights between 14-160 kDa and can be visualized under light microscopy as inclusion bodies (Schnepf et al. 1998). The geometry of these crystals allows a first classification system of Bt strains. In addition, Bt has other insecticidal proteins like Vips that are secreted during its vegetative cycle (Estruch et al. 1996, Schnepf et al. 1998).
Cry genes were classified into 40 families according to their amino acid sequence similarities (Samsonov et al. 1997) and the sequence of more than 160 cry genes are known. The specificity of the toxic effect of the d -endotoxins against certain species makes them environmentally friendly tools for the control of insects that are plagues of important agricultural crops. Cry proteins that could be of particular interest for this study are: Cry3, Cry7 and Cry8 since they have been reported as effective against coleopterans, whereas CryIB, Cry1K and Cry1I have a dual effect against coleopteran and lepidopteran insects (Bravo et al. 1998). In addition, Méndez-López et al. (2003) recently reported that Bacillus thuringiensis serovar israeliensis is toxic to H. hampei.
The identification of Bt cry genes by PCR has proven to be a very useful method for strain characterization and selection (Porcar and Juárez-Pérez, 2003). In addition Carozzi et al. (1991) found correspondence of toxicity with the amplification of particular cry gene profiles, introducing PCR as a tool to predict B. thuringiensis insecticidal activity.
B. thuringiensis-derived bioinsecticides are used in conventional and organic farming (Gustafson et al. 1997), representing 90% of the formulated products that are available in the market. In addition, B. thuringiensis cry genes have allowed the development of transgenic plants that expresse d -endotoxins reducing environmental damage caused by synthetic insecticides and diminishing production costs. The development of new B. thuringiensis-derived bioinsecticides and the manipulation of cry genes for plant genetic engineering depend on the availability of d -endotoxins that differ in insecticidal activity. To date, there are several genetically engineered B. thuringiensis- plants, including coffee modified with the cry1A gene, to confer resistance against Perileucoptera coffeella (Hatanaka et al. 1999). Thus, there is great interest in the isolation of new B. thuringiensis strains to perform wide screening programs with the purpose of searching for new biological activities against insects of economic importance in agriculture (Kim et al. 1998).
Since it has been propose that B. thuringiensis strains coevolved with their insect hosts (Apoyolo et al. 1995), a more directed approach is to isolate B. thuringiensis strains from coffee plantations infected with H. hampei. The main purpose of this research was to isolate B. thuringiensis from environmental samples obtained from coffee plantations infested with the H. hampei and to characterize them by light microscopy, SDS-PAGE, and PCR using specific primers for the cry, cyt and vip gene families.
Materials and methods
Sample collection: 221 samples of soil, leaf litter, fresh foliage and coffee beans from six H. hampei infested coffee plantations were analyzed for the presence of B. thuringiensis. The samples were dried at 50ºC for 24 hr and later stored at room temperature. In addition, 420 groups of 10-15 insects obtained from the dissection of infested coffee beans and insect traps were analyzed. In addition, 23 individual dead larvae found in infested fruits were also evaluated. The insects were stored at 4ºC in 1.5 ml Eppendorf tubes containing 70% ethanol.
Isolation of B. thuringiensis: Bacteria were isolated using the protocol described by Travers et al. (1987). For the isolation from insects, groups of 10-15 adults or individual larvae were homogenized with a sterile pestle in Eppendorf tubes contained LB medium and incubated overnight at 30°C. An aliquot of 200µl of the culture was transferred to Petri dishes with LB-agar medium and incubated at 30°C for 18-24 h. Sporulated colonies with typical B. thuringiensis morphology were preserved on filter paper at room temperature.
Light and scanning electron microscopy: The cell suspensions were fixed to a micro slide by heating, and stained with Coomassie blue for 2 min and observed under a light microscope (Nikon Eclipse E-200). For scanning electron microscopy, diluted spore-crystal suspensions were placed on aluminum stubs and air-dried. Samples were coated with gold in a Fullam EMS-76M Evaporator for 5 min, examined and photographed with a Hitachi S-2360N at 10-15 kv and observed at magnifications between 2000 and 24000X.
Polyacrylamide gel electrophoresis: Spore-crystal suspensions were used for protein analyses in 10% SDS-polyacrylamide gels. Molecular weight markers (BIO-RAD 161-0304) were used to determine the molecular weight of the proteins.
DNA extractions: The protocol for DNA extraction described by Chen and Kuo (1993) was used and DNA concentrations were estimated by fluorometry at 280 nm (Quantech fluorometer, model FM1 109535).
PCR analysis: The following primers were used: cry1 family (cry1Aa, cry1Ab, cry1Ac, cry1Ad, cry1B, cry1C, cry1D, cry1E and cry1F), cry3-cry7, cry5-cry10, cry8, cry11, cry12, cry14, cyt (Bravo et al. 1998) vip1, vip2 and vip3 (Bravo 2003, personal communication) and cry2 (Ben-Dov et al. 1997). PCR conditions were used as follows: 25 mM MgCl2 , 10mM dNTPs, 20 µM of each primer and 2,5 U Taq polymerase. In addition, 1µl of DNA at a concentration of 25 ng/µl was used. The PCR program was: one denaturing cycle of 2 min at 95°C, 30 cycles of 1 min at 95°C, 1 min at 48-54°C, 1 min at 72°C, and a final extension cycle of 5 min at 72°C. PCR products were analyzed by gel electrophoresis in 1% agarose gels.
Results
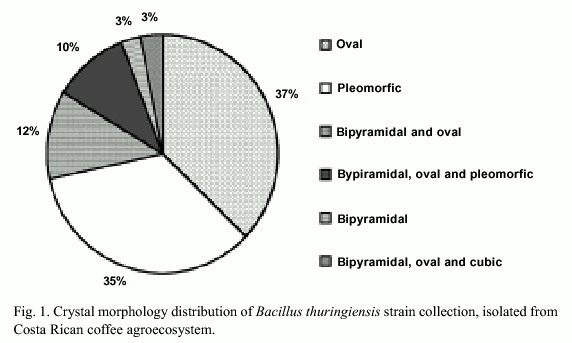
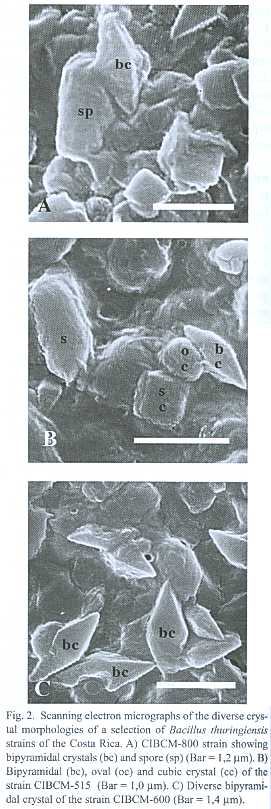
Three hundred strains out of a total of 641 environmental samples examined showed colonial morphology similar to Bacillus thuringiensis. Light microscopy demonstrated that 202 strains presented diverse crystal morphologies: pleomorphic (35%), oval (37%), bipyramidal (3%), bipyramidal and oval (12%), bipyramidal, oval and pleomorphic (10%), bipyramidal, oval and cubic (3%) (Fig. 1). Crystal diversity was confirmed using scanning electron microscopy, showing that some strains presented the crystal adhered to the spore (Fig. 2).
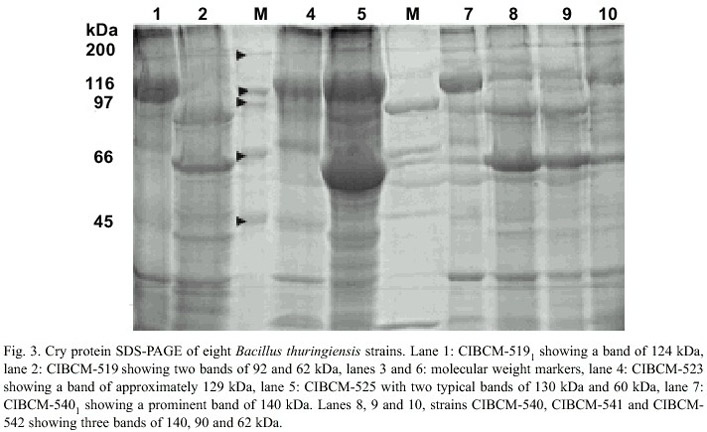
Strains that presented crystals were later analyzed by SDS-PAGE to estimate the molecular weight of the Cry proteins. B. thuringiensis strains presented d -endotoxins with MWs between 20 and 145 kDa but the most common pattern was composed of proteins with MWs between 60-140 kDa (Fig. 3). Five strains presented d -endotoxin from 20 to 40 kDa, six from 40 to 50 kDa, seven from 50 to 60 kDa, 19 from 60 to 70 kDa, 29 from 70 to 100 kDa and 39 from 100-145 kDa.
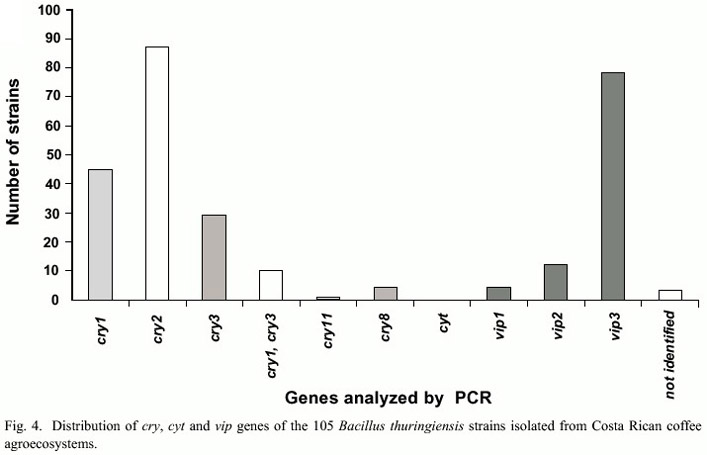
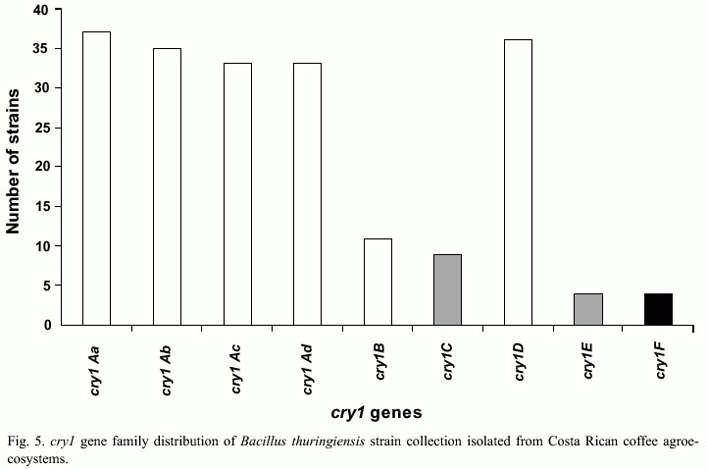
PCR analyses showed that from a total of 105 strains, 82 had the cry2 gene, 45 cry1, 29 cry3-cry7 and 78 the vip3 gene. No amplification was observed for the cry5, cry14, and cry12 and cyt genes. Only one strain amplified with cry11 (Fig. 4). The 45 strains that amplified with the general primer for cry1 family were further characterized with specific primers for this subfamily. It was observed that 36 strains presented cry1D, 37 cry1Aa, 35 cry1Ab and 33 cry1Ac genes, in contrast to other genes that were less frequent, such as cry1E (4 strains) or cry1F genes (4 strains) (Fig. 5). A total of 43 different genetic profiles were found, mainly due to the combination of cry1A genes with other cry and vip genes. It was also frequent to find strains that presented combinations of the cry1, cry2 and cry3-cry7genes. In addition two strains presented the genes cry1, cry3-cry7 and cry8. Ten strains only amplified with the vip3 primers and three did not amplified with any of the primers used.
Discussion
A collection of 202 B. thuringiensis strains were isolated from Costa Rican coffee plantations infested with H. hampei. These strains were analyzed by light microscopy, providing evidence that they produce parasporal crystals. The great diversity observed in crystal morphology for a given strain and between strains could be related to the presence of different and novel d -endotoxins (Fig 2), particularly for those strains that did not amplified with any of the primers tested. These strains may contain insecticidal proteins with specificity towards other insect orders.
It is interesting to notice that while most of the B. thuringiensis strains produce free crystals in the mother cell compartment, it was observed that some presented the crystal adhered to the spore. It is well known that the spore and crystal are liberated to the environment by the action of lytic enzymes that destroy the membrane that contains them (Bechtel & Bulla 1976). However, the absence of this step in some strains of this collection is similar to the one observed in Bacillus sphaericus, were the binary protein crystal is adhered to the spore and covered by the exosporium membrane.
The diversity of molecular weights of the 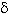
The identification of cry, cyt and vip genes in the B. thuringiensis collection is important, since the biological activity of several Cry proteins is known. This allows the selection of strains to be used in the bioassays for specific insect species (Samsonov et al. 1997). The abundance of strains containing vip3 genes (74%) is a novel finding, if compared to the 15% previously reported by Estruch and collaborators (1996). Vip3 protein has been reported to be toxic for lepidopterans. On the other hand, the genes that code for the Vip1 and Vip2 proteins described as toxic to coleopterans, were found in lower frequencies, but always associated to the vip3 gene.
The most common cry genes found in nature are those within the cry1 subfamily (Porcar and Juárez-Pérez 2003). The most abundant gene found in our collection was cry2, and the second most abundant was the cry1 gene family. Several reports show a high frequency of certain cry1 gene combinations, for example the linkage cry1C-cry1D (Bravo et al. 1998, Ferrandis et al. 1999). The cry1C-cry1D linkage may be explained by their common location on the same replicon (Sanchis et al. 1988). However, in this study cry1D gene was found alone at a relatively high frequency (24 strains) and only 7 strains presented such linkage. The absence of cry1C may be the result of a deletion or a negative selection of cry1C from an ancestral cry1C-cry1D linkage (Ferrandis et al. 1999). One interesting observation was the high frequency of cry1D in combination with those of the subfamily cry1A. The opposite case was observed with the cry1E and cry1F genes, since only eight strains contained these genes. It is also known that cry1B is found at low frequency (Porcar and Juárez-Pérez), however eleven strains of this collection contained this gene; this could be relevant for the purpose of this study because of the known activity of cry1B against coleopterans.
Strains with the cry3-cry7 and cry8 genes are also important in our collection due to their activity against coleopterans (Bernhard et al. 1986). The abundance of these genes in this collection is similar to their prevalence in the collection reported by Bravo et al. (1998). In total, there are 40 strains with toxic potential for coleopterans: 36 strains containing the cry 3-cry7 genes and four strains with cry 8 genes, in addition to eleven strains that have the cry1B genes. However, the strains that did not amplified with cry3, cry7, cry8 and cry1B genes, should be tested in bioassays with H. hampei because they might contain other genes of interest. It is important to notice that even though no B. thuringiensis israelensis-like strains were isolated from coffee plantations, there are several Bti strains isolated from Costa Rican National Parks (Arrieta 2003) that could be tested against the coffee berry borer since this strain was recently reported toxic to H. hampei (Méndez-Lopéz et al. 2003). In conclusion, CIBCM-B. thuringiensis collections might contain strains with potential activities against H. hampei as well as other Cry proteins active against other insect species, therefore, they will be tested in bioassays against specific insect pests in the future.
Acknowledgments
We want to acknowledge the Fundación para la Cooperación Costa Rica-Estados Unidos de América (CR.USA) and the Programa de Cooperación del Gobierno de México for the financial support of the project. We express our acknowledgment to CYTED for supporting the Bt-network of which our laboratory is part. We also thank Alejandra Bravo from the Instituto de Biotecnología de la Universidad Autonoma de México, for critically reading the manuscript.
Resumen
En el año 2000 se reportó por primera vez la principal plaga del cafeto, conocida como broca (Hypothenemus hampei Ferrari) (Coleoptera: Scolitidae) en plantaciones de este cultivo en Costa Rica. Debido al impacto que esta plaga tiene en la economía del país, surgió la necesidad de encontrar una alternativa para el control biológico de esta plaga, basadas en la bacteria entomopatógena Bacillus thuringiensis. El objetivo de este trabajo fue aislar y caracterizar cepas de Bacillus thuringiensis a partir de plantaciones de café infestadas con H. hampei. Se aislaton 202 cepas que se analizaron mediante la morfología del cristal, SDS-PAGE de las d -endotoxinas, mientras que 105 cepas se evaluaron mediante PCR para determinar la presencia de genes cry, cyt y vip. La mayoría de las cepas presentaron diversas morfologías del cristal: pleomórficos (35%), ovalados (37%), bipiramidales (3%), bipiramidales y ovalados (12%), bipiramidales, ovalados y pleomórficos (10%) y bipiramidales, ovalados y cúbicos (3%). El análisis electorforético de las proteínas mostró que 5 cepas contenían d -endotoxinas con pesos moleculares entre los 20 y 40 kDa, 6 entre los 40 y 50 kDa, 7 entre los 50 y 60 kDa, 19 cepas entre los 60 y 70 kDa, 29 entre los 70 y 100 kDa y 39 cepas entre los 100 y 145 kDa. Los análisis mediante PCR mostró que la colección presenta una gran diversidad de genes cry, observándose varios genes por cepa: 78 cepas presentaron el gen vip3, 82 el gen cry2, 45 el gen cry1 y 29 cepas los genes cry3 y cry7. Un total de 13 cepas no amplificaron con los iniciadores cry1, cry2, cry3-7, cry5, cry11cry12 y cry14. Se encontraron 43 perfiles genéticos diferentes, detectándose principalmente la combinación de genes cry1A con otros genes cry o vip. La caracterización genética de esta colección provee información importante para la selección de cepas de Bacillus thuringiensis que se evaluarán mediante bioensayos contra Hypothenemus hampei u otras plagas de importancia económica.
References
Apoyolo, C.I., L. Drif, J.M.Vassal, H. Debarjac, J.P. Bossy, F. Leclant & R. Frutos. 1995. Isolation of multiple subspecies of Bacillus thuringiensis from a population of the European sunflower moth, Homoeosoma nebulella. Appl. Environ. Microbiol. 61: 4343-4347. [ Links ]
Arrieta, G. 2003. Caracterización molecular de las d -endotoxinas de una colección de Bacillus thuringiensis aislada a partir de ecosistemas silvestres y agrícolas de Costa Rica. Tesis de Maestría, Universidad de Costa Rica, San José, Costa Rica. 110 p. [ Links ]
Bechtel, D. B. & L.A. Bulla. 1976. Electron microscopic study of sporulation and parasporal formation in Bacillus thuringiensis. J. Bacteriol. 127: 1472-1481. [ Links ]
Ben-Dov, E., A. Zaritsky, E. Dahan, Z. Barak, R. Sinai, R. Manasherob, A. Khamraev, E. Troitskaya, A. Dubitsky, N. Berezina & Y. Margalith. 1997. Extended screening by PCR for seven cry-group genes from field-collected strains of Bacillus thuringiensis. Appl. Environ. Microbiol. 63: 4883-4890. [ Links ]
Bernhard, K. 1986. Studies on the delta-endotoxin of Bacillus thuringiensis var. Tenebrionis. FEMS Microbiol. Lett. 33: 261-265. [ Links ]
Bernhard, K., Jarret P., M. Meadows, J. Butt, D.J. Ellis, G.M. Roberts, S. Pauli, P. Rogers & H.D. Burges. 1997. Natural isolates of Bacillus thuringiensis: worldwide distribution, characterization and activity against insect pest. J. Invertebr. Pathol. 70: 59-68. [ Links ]
Brun, L.O., C. Marcillaud, V. Gaudichon & D.M. Suckling. 1989. Endosulfan resistance in Hypothenemus hampei (Coleoptera: Scolytidae) in New Caledonia. J. Econ. Entomol. 82: 1312-1316. [ Links ]
Bravo, A., S. Sarabia, L. López, H. Ontiveros, C. Abarca, A. Ortiz, M. Ortiz, L. Lina, J. Villalobos, G. Peña, V. Noez, M. Soberón & R. Quintero. 1998. Characterization of cry genes in a Mexican Bacillus thuringiensis strain collection. Appl. Environ. Microbiol. 64: 4965- 4972. [ Links ]
Carozzi, N. B., V.C. Kramer, G. W. Warren, S. Evola & M.G. Koziel. 1991. Prediction of insecticidal activity of Bacillus thuringiensis strains by polymerase chain reaction product profiles. Appl. Environ. Microbiol. 57: 3057-3061. [ Links ]
Chen W. & T. Kuo. 1993. A simple and rapid method for the preparation of gram-negative bacterial genomic DNA. Nucleic Acid Research 21: 2260. [ Links ]
Estruch, J.J., G.W. Warren, M.A. Mullins, G.J. Nye, J.A. Craig & M.G. Koziel. 1996. Vip3A, a novel Bacillus thuringiensis vegetative insecticidal protein with a wide spectum of activities against lepidopteran insects. Proc. Natl. Acad. Sci. 93: 5389-5394. [ Links ]
Ferrandis, M. D., V. M. Juárez-Pérez, R. Frutos, Y. Bel & J. Ferré. 1999. Distribution of cry1, cryII and cryV genes within Bacillus thuringiensis isolates from Spain. Syst. Appl. Microbiol. 22: 179-185. [ Links ]
Gustafson, M.E., R.A. Clayton, P.B. Lavrik, G.V. Jonhson, R.M. Leimgruber, S.R. Sims & D.E. Bartnicki. 1997. Large-scale production and characterization of Bacillus thuringiensis subsp. tenebrionis insecticidal protein from Escherichia coli. Appl. Microbiol. Biotech. 47: 255-261. [ Links ]
Hatanaka, T., Y. Choi, T. Kusano & H. Sano. 1999. Transgenic plants of coffee Coffea arabica from embryogenic callus via Agrobacterium tumefaciens-mediated transformation. Plant Cell Reports 19: 106-110. [ Links ]
Kim, H.S., D.W. Lee, S.D. Woo, Y.M. Yu & S.K. Kang. 1998. Distribution, serological identification, and PCR analysis of Bacillus thuringiensis isolated from soils of Korea. Curr. Microbiol. 37: 195-20. [ Links ]
Le Pelley, R.H. 1968. Pest of coffee. London, Longman. 590 p. [ Links ]
Méndez-López, I., R. Basurto-Ríos & J. Ibarra. 2003. Bacillus thuringiensis serovar israelensis is highly toxic to the coffee berry borer, Hypothenemus hampei Ferr. (Coleoptera: Scolytidae). FEMS Microbiol. Lett. 11131: 1-5. [ Links ]
Porcar, M. & V. Juárez-Pérez. 2003. PCR-based identification of Bacillus thuringiensis pesticidal crystal genes. FEMS Microbiol. Rev. 26: 419-432. [ Links ]
Samsonov, P., R.I. Padrón, C. Pardo, J. Cabrera & G.A. De al Riva. 1997. Bacillus thuringiensis from biodiversity to biotechnology. J. Ind. Microbiol. Biotech. 19: 202-219. [ Links ]
Sanchis, V., D. Lereclus, G. Menou, J. Chaufaux & M.M. Lecadet. 1988. Multiplicity of d -endotoxin genes with different specificities in Bacillus thuringiensis aizawai. Mol. Microbiol. 2: 393-404. [ Links ]
Schnepf, H.E., N. Crickmore, N. Van Rie, J. Dereclus, J. Baum, J. Feitelson, D.R. Zeigler & D.H. Dean. 1998. Bacillus thuringiensis and its pesticidal crystal proteins. Microbiol. Mol. Biol. Rev. 62: 775-806. [ Links ]
Travers, R. S., P.A.W. Martin & C.F. Reichelderfer. 1987. Selective process for efficient isolation of soil Bacillus spp. Appl. Environ. Microbiol. 53: 1263-1266. [ Links ]