Services on Demand
Journal
Article
Indicators
-
 Cited by SciELO
Cited by SciELO -
 Access statistics
Access statistics
Related links
-
 Similars in
SciELO
Similars in
SciELO  uBio
uBio
Share
Revista de Biología Tropical
On-line version ISSN 0034-7744Print version ISSN 0034-7744
Rev. biol. trop vol.52 n.3 San José Sep. 2004
Interethnic variability and admixture in Latin America - social implications
Francisco M. Salzano
Departamento de Genética, Instituto de Biociências, Universidade Federal do Rio Grande do Sul, Caixa Postal 15053, 91501-970 Porto Alegre, RS, Brazil; francisco.salzano@ufrgs.br
Received 31-X-2003. Corrected 28-XI-2003. Accepted 28-XI-2003.
Abstract
Past and present attempts to classify and characterize the human biological variability are examined, considering the race concept, ethnic identification problems, assortative mating based on ethnicity, and historical genetics. In relation to the latter, a review is made of the methods presently available for admixture quantification and of previous studies aimed at the characterization of the parental continental contributions to Latin American populations, with emphasis in global evaluations of the Costa Rican and Brazilian gene pools. Finally, the question of racism and discrimination is considered, including the relation between human rights and affirmative actions. The right to equal opportunity should be strictly respected. Biological inequality has nothing to do with the ethical principle that someones position in a given society should be an accurate reflection of her/his individual ability. Rev. Biol. Trop. 52(3): 405-415. Epub 2004 Dic 15.
Key words: Race, interethnic variability, assortative mating, racism, ethnic discrimination, affirmative policies.
Palabras clave: Raza, variabilidad interétnica, matrimonio selectivo, racismo, discriminación racial, políticas afirmativas.
Our species is genetically quite variable. Taking advantage of the availability of public genome databases, a map was built of not less than 1.42 million single nucleotide polymorphisms (SNPs, that is, variants involving a single point in the DNA). This nucleotide diversity varies greatly across the genome, but it is estimated that one SNP occurs at every 1.9 kilobases of DNA (The International SNP Map Working Group 2001). Most recent estimates indicate that the number of SNPs in our genome well exceeds two million (Marth et al. 2003). These are large figures, but our closest biological relatives, chimpanzees and bonobos (Pan troglodytes and P. paniscus), are even more variable. Estimates have been made that for mitochondrial DNA these two species show 2- to 4-fold higher levels of diversity, and for the Y-chromosome 6-7 times more variation. For the X-chromosome and autosomal loci the situation is not so clear, but in general higher levels of nucleotide variation were found in Pan (Stone et al. 2002).
The molecular methods which can be used to evaluate our diversity are diverse. Examples are those that determine restriction fragment length polymorphisms (RFLPs) and short tandem repeat polymorphisms (STRPs), besides the SNPs cited above, which can be assessed through sequencing. The changes involve not only single nucleotide changes, but also deletions, insertions, as well as more complex rearrangements.
The picture that is emerging is that this variability is structured, as was verified by Rosenberg et al. (2002), after studying 377 autosomal microsatellite (STR) loci in 1056 individuals from 52 world populations. Despite the fact that within-population differences among individuals account for 93% to 95% of the variation (and similar values have been obtained for blood group, protein, and other DNA markers; Cavalli-Sforza and Feldman 2003), without using prior information about the origins of individuals clusters have been identified corresponding to the major world geographic regions. Similarly, Barbujani and Romualdi (2001) verified that using more than 20 loci, by discriminant analyses, 80% of the genotypes (60% for the Y-chromosome) could be attributed to the right continent. Bamshad et al. (2003) assayed 100 Alu insertion polymorphisms in a heterogeneous collection of 565 individuals, 200 of whom were also typed for 60 microsatellites. Correct assignment to the continent of origin (Africa, Asia, or Europe) reached 99% to 100% when more than 100 loci were used.
Due to different evolutionary histories, the pattern of recombination found in the DNA of peoples from different continents is diverse. Gabriel et al. (2002) characterized haplotype patterns across 51 autosomal regions (spanning 13 megabases) in samples from Africa, Europe, and Asia. They found that half of the human genome exists in blocks of 22 kilobases (kb) or larger in African or African-derived populations, while these blocks are much larger (44 kb or larger) in European and Asian groups.
To evaluate the significance of these findings in social terms, it is necessary to go back to the past, examining how these intercontinental differences had been considered and interpreted, returning afterwards to our time.
The race concept
Inhabitants of different continents can be usually distinguished morphologically, and George Louis Leclerc de Buffon (1707-1788) was the first scholar who applied the term race to different varieties of Homo sapiens (Comas 1966). Initially the static idea of "racial types" (an ideal abstraction of a set of characteristics that would be inherited "en bloc" and therefore could be identified individually) was established. This concept was changed in the first half of the 20th century, with the incorporation of population genetics principles in biological thought. It was then emphasized that race (the taxonomic unit immediately below the species) is a population unit (review in Salzano 1997).
Templeton (1999) questioned the occurrence of human races. According to him these entities cannot be considered as distinct evolutionary lineages, since genetic differentiation alone is insufficient to define a subspecies (race), an additional requirement being historical continuity. Recent admixture would provide sufficient genetic contact to make all of humanity a single lineage sharing a common evolutionary fate.
Another point to be emphasized is that race is also a social concept, and biologically different persons may also have different cultures. Cultural units can be characterized as ethnic groups (from the Greek ethnos, meaning nation). An ethnic group can be a nation, or groups defined by language, social customs or religion. Since the term is intentionally vague Ashley Montagu (see, for instance, Montagu 1960) was one of the first scholars to suggest that the word race should be substituted for ethnic group. This suggestion has been followed in our most recent works.
Independently of names (according to Garn 1971 the term could be ecar, which is simply race spelled backwards) the question remains, is that unit a biological entity? The studies mentioned in the introduction clearly indicate that continental groups ("races") can be clearly distinguished molecularly.
Classification problems
Morphological features, such as skin color, type and color of the hair, nose and lip characteristics are often used by people for self-classification or the categorization of others. In certain communities, however, the descent rule prevails. If a person has an ancestor, no matter how remote, from a given continent, she/he would be classified as derived from that region independently of physical features.
In communities where interethnic marriages are rigidly controlled by customs or law, and segregation policies are enforced, such classifications would be relatively easy to apply. This situation, however, is not generally true in Latin America, where interethnic crosses are widespread. Therefore, complex classifications such as the 16 categories developed in 18th century Spanish America (which included bizarre terms like castizo, morisco, cambujo and ahí te estás) were never socially or legally enforced (Wagley 1971).
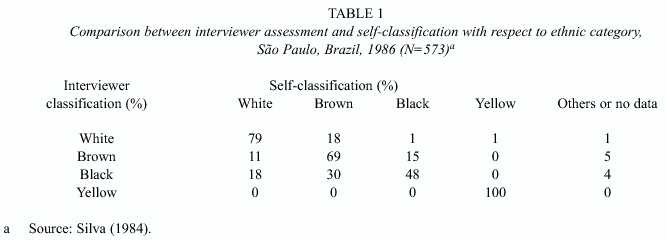
In Brazil, traditionally, the census officers used four categories of skin color (branco, pardo, preto, amarelo, or white, brown, black, yellow); but when the National Research on Sampling by Domicile offered an open question for ethnic self-classification not less than 190 different denominations appeared (PNAD 1976). Matching problems can occur even with the first, rough classification. Table 1 shows the proportion of agreement between the interviewer classification and self-classification in a São Paulo, Brazil, survey conducted in 1986. As can be seen, there was only 79% of agreement in the classification of whites, and the concordance in the brown and black categories was even lower (respectively 69% and 48%).
Assortative mating
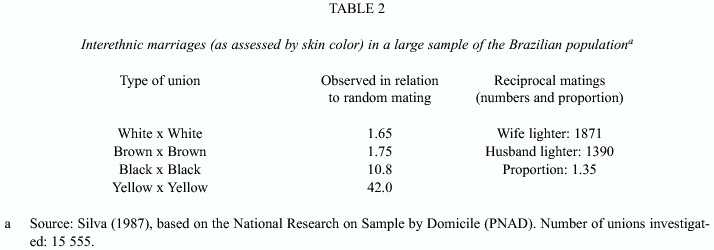
Mating does not occur at random in relation to ethnicity, since persons from different ethnic groups may differ not only in morphology, but also in social class, culture, and other variables. Table 2 presents information about such unions in a large sample of the Brazilian population. There are about 11x more black x black unions and 42x more yellow x yellow unions than those expected by chance. Ethnic endogamy (as assessed by color) is clearly a prevalent event. The direction of the interethnic gene flow also differs according to the sex of the spouses; there are 1.35x more unions in the situation when the wife has a lighter skin than vice versa. Men, if they can choose, marry women with lighter skin. This may be a reflection of the higher social status generally associated with "whiteness". But the situation was different in the past, Colonial times, as will be shown by the genetic data to be presented in later sections.
Historical genetics – admixture quantification
If a given genetic variant is present in one ethnic group only and appears in a mixed population, then it can provide a direct assessment of the historical, accumulated contribution of that ethnic group to the admixed one. Szathmáry and Reed (1978) provided details as how to deal with this type of data. The next simplest case is the one in which just two parental and one descendant population are considered. Allele frequency data can then be used in a model of simple dilution when just one system (or a particularly informative one) is available (Bernstein 1931; Ottensooser 1944). An extension of this method was devised by Ottensooser (1962) that could deal with three parental stocks.
More complex methodologies, that require electronic computation, were afterwards developed. Three much used methods are those of maximum likelihood (Krieger et al. 1965, Wang 2003); weighted least squares (Roberts and Hiorns 1962, 1965, Elston 1971; Long and Smouse 1983, Long 1991, Long et al. 1991) and gene identity (Chakraborty 1975). The method of weighted averages, developed by Cavalli-Sforza and Bodmer (1971) has the limitation that it cannot be used for multiallelic loci (Long and Smouse 1983). Bertorelle and Excoffier (1998) and Dupanloup and Bertorelle (2001) developed a method that is based on a coalescent approach, which explicitly takes into account molecular information as well as gene frequencies. Chikhi et al. (2001) devised a likelihood-based approach using a Markov chain Monte Carlo methodology that also takes into account drift since the admixture event, variation caused by sampling, and uncertainty in the estimates of the ancestral allele frequencies. Their method, however, can only be used in cases in which just two parental populations are involved. Early in these studies it was realized that the best ethnicity estimators are those with large allele frequency differences between the parental groups, and Shriver et al. (1997) provided a list of such alleles; they and McKeighe et al. (2000) also devised a Bayesian log-likelihood analysis for individual and population ethnic-affiliation estimation that was afterwards applied by Parra et al. (2003) to Brazilian data. Parra et al. (1998) and Collins-Schramm et al. (2002) also used alleles with wide allele differences between parental groups (labeled by them as "population- specific" or "ethnic-difference markers") in their analyses.
Relethford (2002) remarked that since the apportionment of genetic diversity in skin color is atypical (only 9% of within-population differences, against the 93%-95% cited above for other markers) it could not be used for purposes of classification. In agreement with that Parra et al. (2003) could find no clear association between individual ethnic evaluations based on 10 population-specific alleles and skin pigmentation coupled with other morphological features, subjectively evaluated, in 173 admixed Brazilians. But conversely, Shriver et al. (2003) found significant correlations between estimates of individual ethnic ancestry based on a panel of 34 ancestry informative markers (another denomination for those with large frequency differences between parental populations) and skin pigmentation, as measured by reflectometry, in African-American and African-Caribbean samples.
Historical genetics – ethnic parental contributions in Latin America
Sans (2000) made a review of the admixture studies performed in Latin America during the 20th century. She classified the populations studied in three categories and was able to list estimates in Central (CA) and South (SA) America as follows: (a) urban, CA:8; SA:17; (b) Amerindian, CA:7; SA:17; and (c) African-derived, CA:3; SA:11. Most of these studies included blood group plus protein markers, and the number of systems considered varied widely. It was clear, however, that the process of admixture is universal in this region of the world, the relative contributions of African, European, and Amerindian gene pools varying in accordance with historical circumstances.
As a part of their global evaluation of the Latin American microcosm, Salzano and Bortolini (2002) also investigated the processes and consequences of interethnic gene flow in the area. Their population classification, however, was different from that of Sans (2000). The study included both previous and the calculation of new estimates, whose number can be categorized as follows: (a) European-derived populations, previous estimates, CA:1; SA:17; new: CA:4; SA:5; (b) African-derived, previous estimates, CA:12; SA:18; new: CA:12; SA:4; and (c) Mestizo or unclassified in relation to continental groups, CA:19; SA:50; new: CA:15; SA:10.
Therefore, no less than 63 Central American and 104 South American evaluations were considered, indicating a good number of studies of this subject. The quality of the data, however, varied markedly among surveys; the reader are referred to the indicated publication for details.
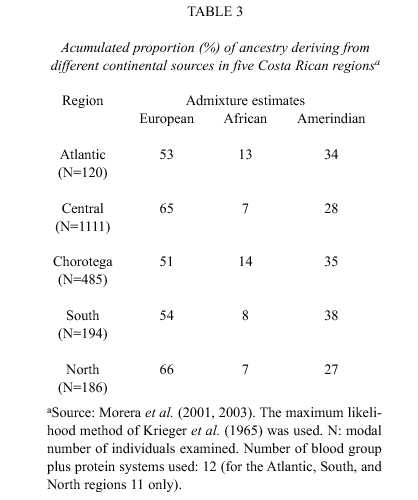
As an example of whole-country evaluations, focus will be given to a Central American (Costa Rica) and a South American (Brazil) populations. Data related to the first is given in Table 3. While in former times it was believed that the people of Costa Rica was essentially of European derivation, the results presented in Table 3 clearly contradict this notion. Only about half (51%-54%) of the genes in the Chorotega, Atlantic, and South regions are estimated to have this derivation, this fraction raising to 65%-66% in the two other regions. The African influence is most marked in the northern Pacific (Chorotega; 14%) and Atlantic (Atlantic; 13%) coasts, while the Amerindian contribution reaches more than one-third of the total (34%-38%) in the above first mentioned (Atlantic, Chorotega, and South) regions.
The Brazilian results are listed in Table 4. Results based on two types of autosomal (blood groups plus proteins; STRPs), as well as mtDNA and Y-chromosome markers, are presented there. Comparing first the two sets of autosome figures, it is seen that the estimated European-derived fraction is markedly larger in the North and Southeast regions when the STRP system is employed (68% vs 52%; 75% vs 52%). This is most probably due to the fact that the sample tested for STRPs was derived from people seeking paternity determinations in a private institution, and most of them had to pay for the tests. Since there are marked differences in socioeconomic status between European-derived and African- or Amerindian-derived persons (a point to be elaborated later), it is possible that this sample presents a higher fraction of persons of European descent. On the other hand, the two sets of results give values which are consistent with the Brazilian history, pointing to a higher Amerindian influence in the North (18%, 48%) and Center-West (11%; 17%) regions. Also according to history, the blood group + protein results suggest higher African contributions in the Northeast (26%) and Southeast (44%) regions.
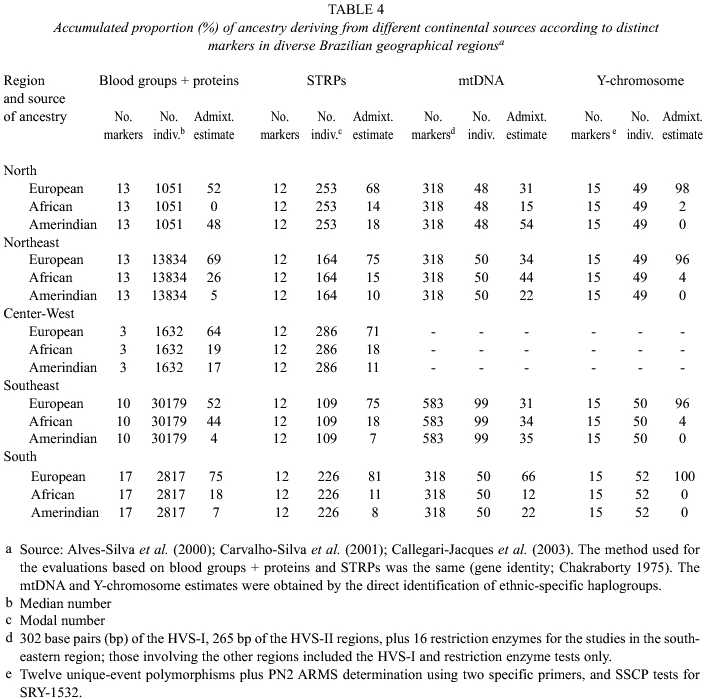
The mtDNA and Y-chromosome data presented in Table 4 emphasize a now well-established fact, that interethnic crosses were asymmetrical in relation to sex in the Brazilian (and Latin American as a whole) past. European males and African or Amerindian females contributed disproportionally more to the process. The Y-chromosomes are over-whelmingly (96%-100%) of European origin, while African (44%) and Amerindian (54%) mtDNA types are especially frequent respectively in the Northeast and North regions.
Ribeiro (1970, 1977) asserted that the history of the conquest of the Americas by Europeans determined the establishment of three types of societies, the transplanted, witness, and new peoples. The first would be formed by descendants of the conquerors who adopted practically without changes the culture and ways of living of the dominant persons. Witness peoples are those who already had a highly developed and complex society, but were forced to witness the new events which occurred in the Conquest without being much affected by them. Finally, the new peoples would have been formed by the complex interaction of the three main groups of founders, Europeans, Africans, and Amerindians. An analogy can be made of this typology for the Latin American genomes. The Y-chromosomes of Latin Americans have been mainly transplanted from outside; their mtDNA, however, received much less outside influences, and can be classified as an witness of the process. This terminology is also adequate, because it indicates the unique possibility of rescuing part of the lost history of extinct aboriginal groups through their permanence in the women of the admixed populations. The nuclear genome, however, due to considerable reshuffling, would be entirely new (Bortolini et al. 2004).
Cerda-Flores et al. (2002a, b) recently reviewed the studies on interethnic admixture in México, providing new estimates of the accumulated proportion of parental groups using D1S80, HLA-DQA1, and 13 STRPs for the northeastern region of that country. The estimates they obtained were of 55%-60% European, 37%-40% Amerindian, and 3%-5% African influence in the present gene pool of northeastern Mexico.
Martínez-Cruzado et al. (2001) were able to rescue mtDNAs from members of the Puerto Rican population that they interpreted to be derived from the Taino Indians, extinct as an independent ethnic unit for centuries now. The Native American maternal contribution to Puerto Ricans was estimated to be 53%; and the prevalence of type 2A genotype of JC virus among them (61%) confirms these findings (Fernandez Cobo et al. 2001).
Similar results have been obtained by Carvajal-Carmona et al. (2000) and Rodas et al. (2003) in the Mestizos of Colombia (respectively 90% of Amerindian mtDNA in Medellín and 78% in Bogotá). As for five African-derived communities of that country Rodas et al. (2003) found 19% of Amerindian mtDNA, while Ribeiro-dos-Santos et al. (2002) observed an even higher prevalence (47%) in Curiaú, a semi-isolated Amazonian Afro-Brazilian community.
Racism and discrimination
The concept of an interethnic system, developed by R.C. Oliveira (see, for instance, Oliveira and Castro Faria 1971) can aptly describe what happens when groups of different ethnic affiliation meet. They are dialectically joined by diametrically opposed, though interdependent interests. Dominance/subordinate relationships generate opposite goals for the two groups, although economically they may depend on each other.
Racism, the assumption of inherent biological superiority of a certain ethnic group in relation to others, and the consequent discrimination against them, generally appears in these situations. Brazil was for a long time considered a "racial paradise", and really the interethnic tensions there did not reach the extremes that were present, for instance, in the United States or South Africa. But the sheer existence of laws about this matter in the country (Afonso Arinos Law, July 3, 1951; Brazilian Constitution of 1988) indicate that racist practices exist there. Presently infractors of these laws are not protected by bail.
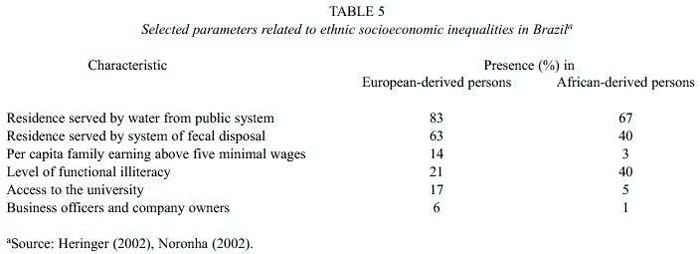
Table 5 presents some selected socioeconomic parameters indicating the different conditions in which European-derived and African-derived persons live in Brazil. As is indicated there, the latter are always in a disadvantaged position, presumably derived from historical times (the slavery stigma) and the perpetuation of dominance/subordinate relationships.
Human rights and affirmative actions
The Universal Declaration of Human Rights, adopted by the United Nations (UN) on December 10, 1948 is a milestone in terms of what should be followed in the relationships between persons or between persons and governments. This document was followed, years afterwards, by the Convention on the Elimination of all Forms of Racial Discrimination, also issued by UN on December 21, 1965. Equally important for the problems discussed here is the United Nations Educational Scientific and Cultural Organization (UNESCO) Declaration on Race and Racial Prejudice of November 27, 1978. A list of the most important documents issued by several organizations related to these matters can be found in Salzano and Hurtado (2004). They generally emphasize individuals rights. The collective rights recognized by international law are relatively few, and they imply the existence of representative collective institutions. It should be remembered that not all traditional custom is by definition good, womens oppression in Moslem societies being just one example. The concept of justice should be universal (Colchester 2002).
The so-called "affirmative actions" are private or public policies against the structural inequalities of groups most vulnerable to discrimination. The term was introduced in the United States especially through two Executive Orders, that of number 10925, issued by president John F. Kennedy in 1961, and that of number 11246, issued by president Lyndon B. Johnson in 1965. The idea was to favor underprivileged persons by extending benefits, such as quotas for admission to employment or universities, to minority groups in general.
In Brazil, the decision to establish affirmative actions occurred after the Third World Conference Against Racism, Xenophobia and Related Intolerance, held in Durban, South Africa, in 2001. Examples are as follows: (a) The Ministry of Justice reserved 20% of executive posts and head positions to African-descendants; (b) Fellowships were created by the Ministry of Foreign Affairs especially for this ethnic category; (c) A quota of 20% for African-derived persons was established in the federal service; (d) The Federal Supreme Court established a 20% quota of African-descendants to be observed in the companies which furnish services to it; and (e) The State University of Rio de Janeiro reserved 40% of its vacancies to persons of this ethnic group (Cesar 2003).
There are doubts about the correctness of these actions. Especially in the case of the 2003 examination entrance to the State University of Rio de Janeiro, no less than 200 judiciary processes were started by persons who considered that they had been unjustly denied access to this institution. In a way, such measures are against the Brazilian constitution of 1988, since they could be considered as racism in backwards. Even in the United States, some persons believe that these programs may actually perpetuate rather than challenge racism (Dominguez 1994). Moreover, the difficulty in the delimitation of ethnic groups or individuals in mixed populations such as those present in Brazil or Latin America as a whole was already emphasized. What level of European, African, or Amerindian ancestry should be considered "significant"? Would self-classification be enough?
Coda
The right to equal opportunity, assured by the Universal Declaration of Human Rights, should be strictly respected. Biological inequality has nothing to do with the ethical principle that someones position in a given society should be an accurate reflection of her/his individual ability.
Acknowledgments
My research is financed by Programa de Apoio a Núcleos de Excelência (PRONEX), Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Fundação de Amparo à Pesquisa do Estado do Rio Grande do Sul (FAPERGS), and Pró-Reitoria de Pesquisa da Universidade Federal do Rio Grande do Sul (PROPESQ/UFRGS).
Resumen
Son examinados los intentos pasados y presentes de clasificar y caracterizar a la variabilidad biológica humana, considerando el concepto de raza, los problemas de identificación étnica, el matrimonio selectivo basado en la etnicidad, y la historia genética. En relación con la última, se hace una revisión de los métodos disponibles actualmente para la cuantificación de la mezcla y de los estudios previos enfocados en la caracterización de la contribuciones parentales de origen continental a las poblaciones Latinoamericanas, con énfasis en las evaluaciones globales de los acervos genéticos de de Brasil y Costa Rica. Finalmente, se considera el tema del racismo y la discriminación, incluyendo la relación entre los derechos humanos y las acciones afirmativas. El derecho a iguales oportunidades debe ser estrictamente respetado. La inequidad biológica no tiene nada que ver con el principio ético de que la posición de cualquiera en una sociedad dada debe ser un reflejo exacto de sus habilidades individuales.
References
Alves-Silva, J., M.S. Santos, P.E.M. Guimarães, A.C.S. Ferreira, H-J. Bandelt, S.D.J. Pena & V.F. Prado. 2000. The ancestry of Brazilian mtDNA lineages. Am. J. Hum. Genet. 67: 444-461. [ Links ]
Bamshad, M.J., S. Wooding, W.S. Watkins, C.T. Ostler, M.A. Batzer & L.B. Jorde. 2003. Human population genetic structure and inference of group membership. Am. J. Hum. Genet. 72: 578-589. [ Links ]
Barbujani, G. & C. Romualdi. 2001. Genetic variation within and among major human groups. Am. J. Hum. Biol. 13: 112-113. [ Links ]
Bernstein, F. 1931. Die geographische Verteilung der Blutgruppen und ihre anthropologische Bedeutung. p. 227-243. In Comitato Italiano per lo Studio dei Problemi della Populazione. Istituto Poligrafico dello Stato, Roma. [ Links ]
Bertorelle, G. & L. Excoffier. 1998. Inferring admixture proportions from molecular data. Mol. Biol. Evol. 15: 1297-1311. [ Links ]
Bortolini, M.C., M.G. Thomas, L. Chikhi, J.A. Aguilar, D. Castro-de-Guerra, F.M. Salzano & A. Ruiz-Linares. 2004. Ribeiros typology, genomes, and Spanish colonialism, as viewed from Gran Canaria and Colombia. Genet. Mol. Biol. 27: 1-8. [ Links ]
Callegari-Jacques, S.M., D. Grattapaglia, F.M. Salzano, S.P. Salamoni, S.G. Crossetti, M.E. Ferreira & M.H. Hutz. 2003. Historical genetics – spatiotemporal analysis on the formation of the Brazilian population. Am. J. Hum. Biol. 15: 824-834. [ Links ]
Cavalli-Sforza, L.L. & W.F. Bodmer. 1971. The Genetics of Human Populations. W.H. Freeman, San Francisco. [ Links ]
Cavalli-Sforza, L.L. & M.W. Feldman. 2003. The application of molecular genetic approaches to the study of human evolution. Nature Genetics 33(suppl.): 266-275. [ Links ]
Carvajal-Carmona, L.G., I.D. Soto, N. Pineda, D. Ortíz-Barrientos, C. Duque, J. Ospina-Duque, M. McCarthy, P. Montoya, V.M. Alvarez, G. Bedoya & A. Ruiz-Linares. 2000. Strong Amerind/White sex bias and a possible Sephardic contribution among the founders of a population in northwest Colombia. Am. J. Hum. Genet. 67: 1287-1295. [ Links ]
Carvalho-Silva, D.R., F.R. Santos, J. Rocha & S.D.J. Pena. 2001. The phylogeography of Brazilian Y-chromosome lineages. Am. J. Hum. Genet. 68: 281-286. [ Links ]
Cerda-Flores, R.M., M.C. Villalobos-Torres, H.A. Barrera-Saldaña, L.M. Cortés-Prieto, L.O. Barajas, F. Rivas, A. Carracedo, Y. Zhong, S.A. Barton & R. Chakraborty. 2002a. Genetic admixture in three Mexican Mestizo populations based on D1S80 and HLA-DQA1 loci. Am. J. Hum. Biol. 14: 257-263. [ Links ]
Cerda-Flores, R.M., B. Budowle, L. Jin, S.A. Barton, R. Deka & R. Chakraborty. 2002b. Maximum likelihood estimates of admixture in northeastern Mexico using 13 short tandem repeat loci. Am. J. Hum. Biol. 14: 429-439. [ Links ]
Cesar, R.C.L. 2003. Ações afirmativas no Brasil: e agora, doutor? Ciênc. Hoje 33(195): 26-32. [ Links ] Chakraborty, R. 1975. Estimation of race admixture: a new method. Am. J. Phys. Anthropol. 42: 507-511. [ Links ]
Chikhi, L., M.W. Bruford & M.A. Beaumont. 2001. Estimation of admixture proportions: a likelihood-based approach using Markov chain Monte Carlo. Genetics 158: 1347-1362. [ Links ]
Colchester, M. 2002. Indigenous rights and the collective conscious. Anthropol. Today 18(1): 1-3. [ Links ]
Collins-Schramm, H.E., C.M. Phillips, D.J. Operario, J.S. Lee, J.L. Weber, R.L. Hanson, W.C. Knowler, R. Cooper, H.Li & M.F. Seldin. 2002. Ethnic-difference markers for use in mapping by admixture linkage disequilibrium. Am. J. Hum. Genet. 70: 737-750. [ Links ]
Comas, J. 1966. Manual de Antropología Física. Universidad Nacional Autónoma de Mexico, Mexico. [ Links ] Domínguez, V.R. 1994. A taste for "the other". Intelectual complicity in racializing practices. Curr. Anthropol. 35: 333-348. [ Links ]
Dupanloup, I. & G. Bertorelle. 2001. Inferring admixture proportions from molecular data: extension to any number of parental populations. Mol. Biol. Evol. 18: 672-675. [ Links ]
Elston, R.C. 1971. The estimation of admixture in racial hybrids. Ann. Hum. Genet. 35: 9-17. [ Links ]
Fernandez Cobo, M., D.V. Jobes, R. Yanagihara, V.R. Nerurkar, Y. Yamamura, C.F. Ryschkewitsch & G.L. Stoner. 2001. Reconstructing population history using JC virus: Amerinds, Spanish, and Africans in the ancestry of modern Puerto Ricans. Hum. Biol. 73: 385-402. [ Links ]
Gabriel, S.B., S.F. Schaffner, H. Nguyen, J.M. Moore, J. Roy, B. Blumenstiel, J. Higgins, M. DeFelice, A. Lochner, M. Faggart, S.N. Lin-Cordero, C. Rotimi, A. Adeyemo, R. Cooper, R. Ward, E.S. Lander, M.J. Daly & D. Altshuler. 2002. The structure of haplotype blocks in the human genome. Science 296: 2225-2229. [ Links ]
Garn, S.M. 1971. Human Races. Charles C Thomas, Springfield. [ Links ] Heringer, R. 2002. Desigualdades raciais no Brasil: síntese de indicadores e desafios no campo das políticas públicas. Cad. Saúde Publ. 18(suppl.): 57-65. [ Links ]
Krieger, H., N.E. Morton, M.P. Mi, E. Azevêdo, A. Freire-Maia & N. Yasuda. 1965. Racial admixture in north-eastern Brasil. Ann. Hum. Genet. 29: 113-125. [ Links ]
Long, J.C. & P.E. Smouse. 1983. Intertribal gene flow between the Yecuana and Yanomama: genetic analysis of an admixed village. Am. J. Phys. Anthropol. 61: 411-422. [ Links ]
Marth, G., G. Schuler, R. Yeh, R. Davenport, R. Agarwala, D. Church, S. Wheelan, J. Baker, M. Ward, M. Kholodov, L. Phan, E. Czabarka, J. Murvai, D. Cutler, S. Wooding, A. Rogers, A. Chakravarti, H.C. Harpending, P-Y. Kwok & S.T. Sherry. 2003. Sequence variations in the public human genome data reflect a bottlenecked population history. Proc. Nat. Acad. Sci., USA 100: 376-381. [ Links ]
Martínez-Cruzado, J.C., G. Toro-Labrador, V. Ho-Fung, M.A. Estévez-Montero, A. Lobaina-Manzanet, D.A. Padovani-Claudio, H. Sánchez-Cruz, P. Ortiz-Bermúdez & A. Sánchez-Crespo. 2001. Mitochondrial DNA analysis reveals substantial Native American ancestry in Puerto Rico. Hum. Biol. 73: 491-511. [ Links ]
McKeigue, P.M., J. Carpenter, E.J. Parra & M.D. Shriver. 2000. Estimation of admixture and detection of link-age in admixed populations by a Bayesian approach: application to African-American populations. Ann. Hum. Genet. 64: 171-186. [ Links ]
Montagu, M.F.A. 1960. An Introduction to Physical Anthropology. Charles C Thomas, Springfield. [ Links ] Morera, B., R. Marin-Rojas & R. Barrantes. 2001 Análisis de varios marcadores genéticos clásicos en la población de Costa Rica. Rev. Biol. Trop. 49: 1237-1252. [ Links ]
Morera, B., R. Barrantes & R. Marin-Rojas. 2003. Gene admixture in the Costa Rican population. Ann. Hum. Genet. 67: 71-80. [ Links ]
Noronha, S. Maioria e minorias no Brasil. Rumos 26(196): 26-33. [ Links ]
Oliveira, R.C. & L. Castro Faria. 1971. Interethnic contact and the study of populations, pp. 41-59. In F.M. Salzano (ed.). The Ongoing Evolution of Latin American Populations. Charles C Thomas, Springfield. [ Links ]
Ottensooser, F. 1944. Cálculo do grau de mistura racial através dos grupos sangüíneos. Rev. Bras. Biol. 4: 531-537. [ Links ]
Ottensooser, F. 1962. Analysis of trihybrid populations. Am. J. Hum. Genet. 14: 278-280. [ Links ]
Parra, E.J., A. Marcini, J. Akey, J. Martinson, M.A. Batzer, R. Cooper, T. Forrester, D.B. Allison, R. Deka, R.E. Ferrell & M.D. Shriver. 1998. Estimating African American admixture proportions by use of population-specific alleles. Am. J. Hum. Genet. 63: 1839-1851. [ Links ]
Parra, F., R.C. Amado, J.R. Lambertucci, J. Rocha, C.M. Antunes & S.D.J. Pena. 2003. Color and genomic ancestry in Brazilians. Proc. Nat. Acad. Sci., USA 100: 177-182. [ Links ]
PNAD. 1976. Pesquisa Nacional por Amostras de Domicilios. Instituto Brasileiro de Geografia e Estatística, Rio de Janeiro. [ Links ]
Relethford, J.H. 2002. Apportionment of global genetic diversity based on craniometrics and skin color. Am. J. Phys. Anthropol. 118: 393-398. [ Links ]
Ribeiro, D. 1970. The culture-historical configurations of the American peoples. Curr. Anthropol. 11: 403-434. [ Links ]
Ribeiro, D. 1977. As Américas e a Civilização. Vozes, Petrópolis. [ Links ]
Ribeiro-dos-Santos, A.K.C., J.A. Pereira, M.F.R. Lobato, B.M. Carvalho, J.F. Guerreiro & S.E.B. Santos. 2002. Dissimilarities in the process of formation of Curiaú, a semi-isolated Afro-Brazilian population of the Amazon region. Am. J. Hum. Biol. 14: 440-447. [ Links ]
Roberts, D.F. & R.W. Hiorns. 1962. The dynamics of racial admixture. Am. J. Hum. Genet. 14: 261-277. [ Links ]
Roberts, D.F. & R.W. Hiorns. 1965. Methods of analysis of the genetic composition of a hybrid population. Hum. Biol. 37: 38-43. [ Links ]
Rodas, C., N. Gelvez & G. Keyeux. 2003. Mitochondrial DNA studies show asymmetrical Amerindian admixture in Afro-Colombian and Mestizo populations. Hum. Biol. 75:13-30. [ Links ]
Rosenberg, N.A., J.K. Pritchard, J.L. Weber, H.M. Cann, K.K. Kidd, L.A. Zhivotovsky & M.W. Feldman. 2002. Genetic structure of human populations. Science 298: 2381-2385. [ Links ]
Salzano, F.M. 1997. Human races: myth, invention, or reality? Interciencia 22: 221-227. [ Links ]
Salzano, F.M. & M.C. Bortolini. 2002. The Evolution and Genetics of Latin American Populations. Cambridge University, Cambridge. [ Links ]
Salzano, F.M. & A.M. Hurtado. 2004. Lost Paradises and the Ethics of Research and Publication. Oxford University, New York. [ Links ]
Sans, M. 2000. Admixture studies in Latin America: from the 20th to the 21st century. Hum. Biol. 72: 155-177. [ Links ]
Shriver, M.D., M.W. Smith, L. Jin, A. Marcini, J.M. Akey, R. Deka & R.E. Ferrell. 1997. Ethnic-affiliation estimation by use of population-specific DNA markers. Am. J. Hum. Genet. 60: 957-964. [ Links ]
Shriver, M.D., E.J. Parra, S. Dios, C. Bonilla, H. Norton, C. Jovel, C. Pfaff, C. Jones, A. Massac, N. Cameron, A. Baron, T. Jackson, G. Argyropoulos, L. Jin, C.J. Hoggart, P.M. McKeigue & R.A. Kittles. 2003. Skin pigmentation, biogeographical ancestry and admixture mapping. Hum. Genet. 112: 387-399. [ Links ]
Silva, N.V. 1987. Distância social e casamento interracial no Brasil. Est. Afro-Asiat. 14: 54-83. [ Links ]
Silva, N.V. 1994. Uma nota sobre "raça social" no Brasil. Est. Afro-Asiat. 26: 67-80. [ Links ]
Stone, A.C., R.C. Griffiths, S.L. Zegura & M.F. Hammer. 2002. High levels of Y-chromosome nucleotide diversity in the genus Pan. Proc. Nat. Acad. Sci., USA 99: 43-48. [ Links ]
Szathmáry, E.J.E. & T.E. Reed. 1978. Calculation of the maximum amount of gene admixture in a hybrid population. Am. J. Phys. Anthropol. 48: 29-34. [ Links ]
Templeton, A.R. 1999. Human races: a genetic and evolutionary perspective. Am. Anthropol. 100: 632-650. [ Links ] The International SNP Map Working Group. 2001. A map of human genome sequence variation containing 1.42 million single nucleotide polymorphisms. Nature 409: 928-933. [ Links ]
Wagley, C. 1971. The formation of the American population, pp. 19-39. In F.M. Salzano (ed.). The Ongoing Evolution of Latin American Populations. Chales C Thomas, Springfield. [ Links ]
Wang, J. 2003. Maximum-likelihood estimation of admixture proportions from genetic data. Genetics 164: 747-765. [ Links ]