Serviços Personalizados
Journal
Artigo
Indicadores
-
 Citado por SciELO
Citado por SciELO -
 Acessos
Acessos
Links relacionados
-
 Similares em
SciELO
Similares em
SciELO
Compartilhar
Acta Médica Costarricense
versão On-line ISSN 0001-6002versão impressa ISSN 0001-6012
Acta méd. costarric vol.63 no.2 San José Abr./Jun. 2021
Original
Viral loads in transplant patients and cytomegalovirus genotyping
1Hospital San Juan de Dios, Laboratorio de Biología Molecular. Costa Rica
2Hospital San Juan de Dios, Servicio de Hematología, San José, Costa Rica.
3Hospital San Juan de Dios, Servicio de Nefrología. Costa Rica
4Universidad de Costa Rica, Escuela de Biología, Laboratorio de Biología Molecular, San José, Costa Rica.
The replacement of human cells, tissues, or organs is used as a therapy to restore or establish the normal function of damaged tissue or organ. 1
Since the first successful transplant was performed, a vast experience in the subject has been generated, developing a series of procedures that depend on the type of transplant to be performed.
When talking about solid organ transplants such as kidney transplants, the organ usually comes from a donor who may be living or also from a dead donor, who is compatible with the recipient or transplanted person.2
In the case of hematopoietic progenitor cell or bone marrow transplants, the criteria become more selective and because of the same complexity, a classification has been established to choose the most appropriate one for the patient. Several diseases related to hematopoietic progenitor cells have been successfully treated by bone marrow allotransplantation.2
Viruses belonging to the Herpesvirus group are those that most frequently infect transplant patients. The importance of Herpesviruses in this population is justified by their ubiquity and their ability to produce latency, which allows the virus to reactivate and begin its replicative cycle in conditions in which there is a decrease in immune surveillance, as occurs in transplants.3
There are factors associated with transplantation based on which decisions are made for the therapeutic management of the patient. Among these factors are the states of neutropenia and immunosuppression triggered by the medications applied for the graft to be successfully achieved and to reduce the possibility of transplant rejection.4 Another factor is the appearance of infections due to the reactivation of previously established latent viruses, even in early childhood.5
The latency of viral infections is characterized by three general properties: viral gene products that promote virus replication in small quantities, cells that harbor the latent genome and are poorly recognized by the immune system, and the persistence of the intact viral genome to initiate a subsequent productive infection, ensuring the spread of viral progeny to new hosts.6 The latency of certain Herpesvirus genomes can be maintained as an episome in dividing cells, as is the case with Epstein- Barr virus (EBV) in B cells or Cytomegalovirus infection in salivary and mammary gland cells. Cytomegalovirus (HCMV) infection is one of the most significant causes of morbidity and mortality in individuals who are affected by episodes of immunosuppression such as transplantation.7 HCMV represents the first cause of primary infection in humans and remains latent throughout life so that reactivation can occur leading to reinfection.6 The seropositivity rate of the general population in developed countries ranges between 30% and 70% and is higher in older and low socioeconomic subjects.8
HCMV glycoprotein B (gB) is a highly immunogenic protein incorporated into the viral envelope, which exerts an essential biological role in the virus-host interaction, as it participates in the entry, propagation, and replication of the virus in different host cells. 9,10. Wild-type HCMV strains can be classified into four major gB genotypic variants (gB 1-4), based on the gB sequence. Each has a tropism for distinct cell lines, leading to differences in pathogenesis and disease severity.
Therefore, genotypic characterization of HCMV strains infecting immunocompromised individuals may contribute to molecular epidemiological studies and the definition of the role of viral genetic variability in clinical expression and prognosis.11
This study aimed to determine the course of viral infections over a period of one year, by measuring the viral loads of Adenovirus, BK virus, Epstein-Barr virus, Cytomegalovirus, and human Herpesvirus 6, in patients undergoing kidney or hematopoietic progenitor cell transplantation, treated at the Hospital San Juan de Dios (HSJD).
METHODS
Patients: Thirty patients older than 12 years with a diagnosis of kidney transplantation (KT) and hematopoietic progenitor cell or bone marrow transplantation (BMT) were studied. Samples from patients of the Nephrology Service and Hematology Service of the HSJD of the Caja Costarricense del Seguro Social (CCSS), during a period of one year (September 2015 to September 2016) were analyzed. The number of transplanted patients in the study was calculated based on the annual historical behavior recorded at the HSJD in previous years (Source: Nephrology and Hematology Service, HSJDCCSS). All patients underwent the informed consent process, according to the procedures and protocol established by the Local Bioethics and Research Committee of the HSJD, CCSS (CLOBI), and the Scientific Ethical Committee of the Tropical Diseases Research Center (CIET), Faculty of Microbiology and Vice-Rectory of Research, University of Costa Rica.
Method: Blood was collected through venipuncture in Vacutainer tubes with EDTA anticuagulant. The blood sample was centrifuged at 2000 g (4400 r.p.m.) for 20 minutes, to separate the plasma which was used in this study. We performed measurements in 10 samples per patient that included: a pre-transplant sample that corresponded to measurement 1 and then 8 samples every fifteen days after the transplant for a period of 4 months (measurements 2 to 9) and the last sample at 6 months after the transplant (measurement 10).
Quantification and detection of virus nucleic acid: Quantification of BK, HCMV, EBV, and HHV6 viruses was performed through real-time polymerase chain reaction (PCR) to determine the viral load of each one. Only in the case of Adenovirus, the detection was performed qualitatively, by end-point PCR.
For HCMV, the ABBOTT technique and equipment were used. The nucleic acid was extracted through electromagnetic particles that trap nucleic acids in the Abbott m2000sp equipment. Subsequently, the reaction mixture was assembled and finally, the HCMV viral load was detected and quantified in the Abbott m2000rt equipment. For the rest of the viruses, a single extraction of nucleic acid was carried out using QIAGEN columns in the QIAcube instrument. The principle of the technique is based on several centrifugation cycles through which the viral genetic material is trapped in the silica gel columns and then the DNA is released employing an elution solution, to be collected in polypropylene tubes. The final volume of nucleic acid extracted was 100 uL. From this extracted DNA, each reaction mixture was assembled according to the virus to be determined and the viral load of each virus was determined by real-time PCR. The real-time PCR reactions were performed in the Rotor-Gene equipment of QIAGEN. In the case of Adenovirus, reagents from SECASE were used to assemble the reaction mixture and nucleic acid amplification was performed by endpoint PCR in a thermocycler from Applied Biosystems. The electrophoretic run of the amplified fragment was then performed on an agarose gel stained with ethidium bromide and visualized through a transilluminator with ultraviolet light. Samples were qualitatively classified as positive or negative. Viral loads of the remaining viruses were reported in terms of copies/mL. Any viral load greater than the lower limit of detection was considered a positive test and the upper and lower limits of detection were determined by commercially available and manufacturer’s recommendations.
Nested PCR and HCMV genotyping: HCMV-positive viral loads were stored in a freezer that maintains samples at -80 °C. For HCMV genotyping purposes, only viral loads greater than 1000 copies/ mL6 were worked with and a nested PCR was performed to verify the presence of viral genetic material using an agarose gel. Nucleic acid extraction was performed from the samples saved and selected for genotyping analysis using QIAGEN brand columns on the QIAcube instrument. For nested PCR, a reaction mix from Thermo Scientific commercial house No. K0171 was used. For the first round of amplification, the external primer oligonucleotides used were gB 1319 (5’TGGAACTGGAACGTTTGGC3’) and gB 1676 (5’TGACGCTGGTTTGGTTGAATG3’). For the second round of nested PCR, primers gB 1319 (5’TGGAACTGGAACGTTTGGC3’) and gB 1604 (5’GAAACGCGCGGCAATCGG3’). were used. All primers were standardized for use at a final concentration of 10 uM. The reaction mixture had a final volume of 50 uL including the reaction solution, each of the primers, DNA from each clinical sample, and nuclease-free water. PCR conditions were as follows: initial denaturation at 95 °C for 3 minutes, 25-40 cycles with denaturation at 95 °C for 30 seconds; hybridization at Tm 55 °C for 30 seconds, and extension at 72 °C for 1 minute, with a final extension at 72 °C for 5-15 minutes.
The 357 bp amplified product from the first round was visualized on a 2% agarose gel stained with ethidium bromide. A nested PCR was performed on all the amplified samples from the first round to obtain an amplified fragment of 285 bp. The conditions and quantities for the second round were the same as for the first round, but with the primers mentioned above. For sequencing, the services of the CIBCM (Centro de Investigación en Biología Celular y Molecular, Universidad de Costa Rica) were contracted, where the samples were analyzed on ABI 3500 equipment. The sequences obtained from the patients were edited to leave only the nucleotide sequence corresponding to codons 441- 511 of the gene for human HCMV glycoprotein B (gB). For this purpose, an alignment of the sequences obtained with the reference sequence Human Herpesvirus 5 strain HANRTR5, complete genome (accession number KY123652.1) was performed using the DNA Baser program (Version 4.36.0.2.). The amino acid sequence was compared by Blast analysis on the NCBI site (https://blast.ncbi.nlm.nih.gov) and a Clustal alignment of the amino acid sequences was performed to compare the amino acid positions in the protein. Clustal alignment was performed using the Clustal Omega tool (http://www.ebi.ac.uk/Tools/msa/clustalo/).
Results
Of the selected sample of 30 patients, 21 underwent KT and 9 underwent BMT and the distribution by sex was 15 men and 15 women, with only one patient under 18 years of age. In addition, one patient had graft rejection on the second day after transplantation and three patients died approximately one and a half months after receiving the graft. We found that 23 patients (77%) tested positive for at least one of the viruses tested (Table 1).
Table 1 Distribution of the 30 patients with kidney transplantation (KT) and hematopoietic progenitor cell or bone marrow transplantation (BMT) according to the type of virus found, time period September 2015 to September 2016, Nephrology Service and Hematology Service, Hospital San Juan de Dios, Caja Costarricense de Seguro Social.
Quantification and detection of virus nucleic acid:
Table 1 shows the viruses found in each patient. A total of 7 patients were found positive for adenovirus, of which 3 corresponded to BMT and 4 to kidney transplantation. BK virus was detected in 5 cases, of which 2 were from KT and 3 from BMT.
The virus most frequently detected was HCMV, with 17 cases corresponding to 11 cases of KT and 6 of BMT. EBV was the least detected virus, with only 2 cases, and finally HHV6 with 3 positive cases.
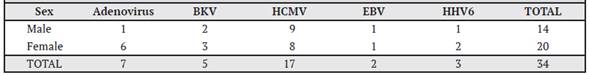
In the Adenovirus-positive patients, one positive qualitative measurement was obtained in measurement 3, two in measurement 4, one in measurement 5, one in measurement 6, one in measurement 7, and one in the post-transplant measurement. Table 2 shows the number of positive patients according to the virus and sex of the patient. In general, there were more viruses present in the female population than in the male population. Adenovirus was present in 6 cases in women and 1 case in men.
Of the 5 positive cases of BK virus, 2 were male and 3 were female. HCMV showed almost equal behavior between men and women, since of the total of 17 positive cases, 9 were male and 8 were female. For Epstein-Barr virus, there was only one female and one male case, while for HHV6 there were 2 cases in females and one in males.
Figure 1A shows the behavior over time of the viral load determinations of the BKV-positive patients. It was observed that in patient 15 the viral load was significantly elevated during measurement 5, then declined and rose again in measurement 9, while in the other patients’ virus was detected in measurements 7 through 9 in the study.
Figure 1B shows the behavior of the viral load over time for Epstein-Barr virus in patients 23 and 24. In both patients, the virus was detected in the fourth measurement.
The only three cases that were positive for HHV6 are shown in Figure 1C, where similar behavior is observed. Patient 3, virus was detected in measurement 2, while patients 13 and 19 virus was detected during measurement three of the study.
Figure 1D shows the behavior over time of the viral loads of HCMV-positive patients. This was the virus that reported the most positive cases, with high viral load values.
Patient 15 and patient 23 were the only patients in whom multiple viruses were detected simultaneously, as shown in Figure 2.
Nested PCR and HCMV genotype detection:
Of the 17 HCMV-positive cases, only 13 patients with viral loads greater than 1000 copies/mL were considered. Of these 13 patients, only 11 samples were amplified in the second round of PCR, and 8 of these samples corresponded to KT patients, and the remaining 3 were from BMT patients. The bands obtained were approximately 285 bp in size. Finally, of the 11 samples amplified in the second round of PCR, samples from 10 patients were successfully sequenced and analyzed (Table 3). Only one patient (number 15) exhibited an HCMV genotype 1, while the others showed HCMV genotype 3. No mixed infection was detected in the patients.
Table 2 Distribution of the 30 patients according to viral detections and type of transplant, time period from September 2015 to September 2016, Nephrology Service and Hematology Service, Hospital San Juan de Dios, Caja Costarricense de Seguro Social.
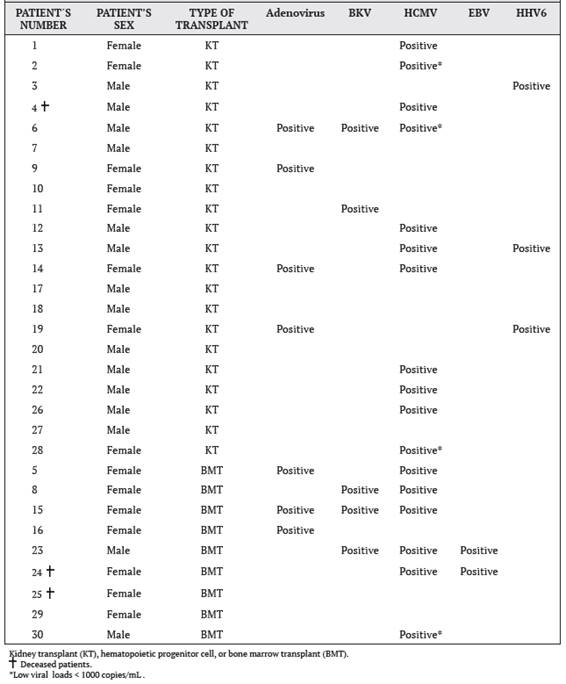
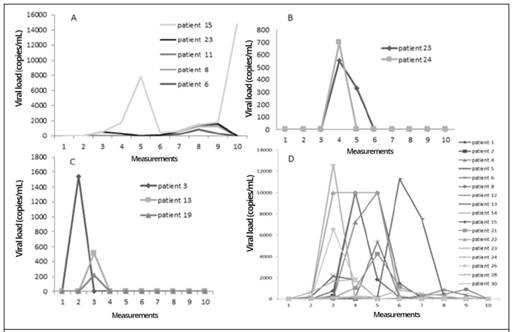
Figure 1 Measurements over time of viral loads in BMT and KT patients, with positive results for four selected viruses: A. BKV, B. EBV, C. HHV6, and D. HCMV. Measurement 1 is pre-transplant, measurements 2-9 were performed every 15 days post-transplant, and measurement 10 was performed at 6 months post-transplant; time period from September 2015 to September 2016, Nephrology Service and Hematology Service, Hospital San Juan de Dios, Caja Costarricense de Seguro Social.
Discussion
For one year, the detection of five viruses was carried out in thirty patients of KT and BMT of the Hospital San Juan de Dios, with the approval of informed consent from each of the patients.
In particular, bone marrow transplant patients, due to the previous medication they receive, suffer from a high depletion of cells, especially lymphocytes, which leads to a favorable environment for the development of many microorganisms, especially viruses. These viral infections can develop as a result of a primoinfection or latency due to past infections and, according to clinical trials, 30% of deaths due to infections after transplantation are due to viruses.15 Concomitant infections with several viruses at the same time have also been observed in this patient population.13
Generally, the activation of common Herpesviruses such as HCMV promotes the reactivation of other latent Herpesviruses such as EBV, BK virus, and Adenovirus. Of all the patients in this study, seven were positive for Adenovirus, i.e. 23%, which is in agreement with other studies indicating that the incidence of Adenovirus in transplant patients is higher than 21%.13,14
Therapeutic management with antivirals causes the viral load to decrease as observed in this study where patients passed several positive measurements but then the viral load drops to undetectable levels by drug administration. It is worth mentioning that these treatments, unfortunately, bring side effects to the patient, especially hematological abnormalities, mainly neutropenia, anemia, and thrombocytopenia.17
For this reason, periodic measurements of viral load in these processes are of great help since only those cases that warrant it will be treated. The high toxicity and the possibility of resistance to these antivirals is an issue that should be taken into account for institutional protocols since it is valid to perform therapeutic management by reducing immunosuppression and observing the patient’s reaction.6,13,16 In addition, the cost for the institution per patient would be lower, since these drugs are expensive.
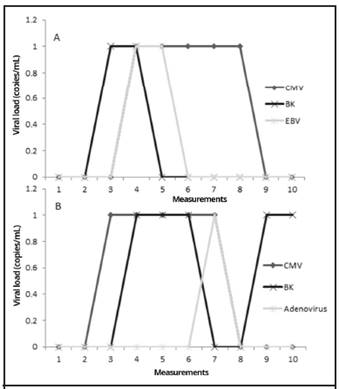
Figure 2 Time-course measurements of viral loads of A. BKV, HCMV, and EBV from BMT patient number 23 and B. BKV, HCMV, and Adenovirus from BMT patient number 15. Measurement 1 is pre-transplant, measurements 2-9 were performed every 15 days post-transplant, and measurement 10 was performed at 6 months post-transplant; time period from September 2015 to September 2016, Nephrology Service and Hematology Service, Hospital San Juan de Dios, Caja Costarricense de Seguro Social.
The study shows a low incidence of EBV and HHV6 viruses in comparison with HCMV, which is curious since according to the literature, the positivity for EBV in transplant patients could reach up to 20%16 and 95% of adults are seropositive for HHV6. 3 This study showed that 2 patients out of 30 were positive for EBV (6%) and only 3 out of 30 were positive for HHV6 (10%).
The study also showed that the virus that reported the most cases was HCMV with 17 positive patients, equivalent to 57% of the total number of patients involved in the project, which makes sense since 70% of the adult population worldwide is seropositive for HCMV.6 Ten HCMV-positive patients underwent HCMV genotyping by sequencing and the most predominant genotype was genotype 3 (gB3), which was present in 9 patients. The remaining patient was the only case of genotype 1 (gB1) and corresponds to a kidney transplant patient. Studies indicate that genotype 1 is not associated with fatal HCMV disease. 18
According to other studies,19 the predominant genotype in Costa Rica was genotype 2 (gB2), however, in a smaller proportion, they also found genotypes 1 and 3. On the other hand, Cunha et al 11 mention that genotypes 2 and 3 are the most frequently found in patients with bone marrow transplants.
Our study is not conclusive, since it has several limitations such as a limited sampling, only plasma samples were used, and the study population refers only to transplant patients. For this reason, it is suggested that in the future, a study with a larger sampling that is more representative of the population in question should be carried out.
Table 3 Multiple alignment of amino acid sequences deduced from the sequence of codons 441-511 of the gene for HCMV glycoprotein B (gB), obtained from blood samples of the 10 HCMV-positive patients and with a viral load greater than 1000 copies/mL, time period from September 2015 to September 2016, Nephrology Service and Hematology Service, Hospital San Juan de Dios, Caja Costarricense de Seguro Social.
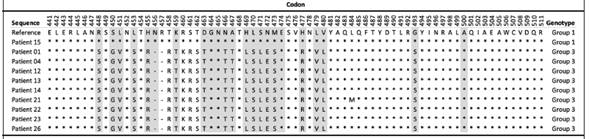

Note: Amino acid positions that differ from the reference sequence used for HCMV (GenBank accession number: KY123652.1) are indicated with a letter symbol, matching positions with an asterisk (* ) ,and absent positions with a dash (-). The HCMV genotype determined for each patient is shown in the last column and the gray shading corresponds to the most relevant positions for determining the HCMV genotype, according to Chou and Dennison (1991).
In conclusion, it can be stated that molecular techniques are an excellent tool for the detection and quantification of viruses and thus the treating physician is able to give better clinical management to the patient suffering from an opportunistic viral infection. In addition, a better prognosis can be achieved without having to use pharmacological therapies that lead to undesirable side effects.20 Acknowledgments and collaborators: To the advisors at the University of Costa Rica: MSc. Sandra Silva De la Fuente, Dr. Rodrigo Mora Rodríguez. Bioinformatics Advisor: Lic. Mario Ulate Solano. To the Direction of the Clinical Laboratory of the Hospital San Juan de Dios: Dr. José Pablo Marín Gómez.
REFERENCES
1. Rouchi H, Mazdeh M. Regenerative Medicine in Organ and Tissue Transplantation: Shortly and Practically Achievable?. Int J Organ Transplant Med. 2015;6(3):93-5 [ Links ]
2. Chinen J, Buckley RH. Transplantation immunology: solid organ and bone marrow. J Allergy Clin Immunol. 2010;125(2): S324-35. [ Links ]
3. Braun DK, Domínguez G, Pellett PE. Human herpesvirus 6. Clin Microbiol Rev. 1997;10(3):521-67. [ Links ]
4. Cervera C, Lumbreras C. Factores de riesgo de la enfermedad por citomegalovirus en el receptor de un trasplante de órgano sólido. Enferm Infecc Microbiol Clin. 2011;29:11-7. [ Links ]
5. Echavarria M, Basilotta N, Aguiar A, Davalos M, Ricarte C, Iotti A, et al . Neuropatía por virus BK post trasplante renal diagnóstico y seguimiento por PCR en tiempo real. Medicina (B Aires). 2007;67(6):719-22. [ Links ]
6. Crough T, Khanna R. Immunobiology of human cytomegalovirus: from bench to bedside. Clin Microbiol Rev. 2009;22(1):76-98. [ Links ]
7. Boeckh M, Boivin G. Quantitation of cytomegalovirus: methodologic aspects and clinical applications. Clin Microbiol Rev. 1998;11(3):533-54. [ Links ]
8. Kanj SS, Sharara AI, Clavien PA, Hamilton JD. Cytomegalovirus infection following liver transplantation: review of the literature. Clin Infect Dis. 1996;22(3):537-49. [ Links ]
9. Arista S, De Grazia S, Giammanco GM, Di Carlo P, Iannitto E. Human cytomegalovirus glycoproteinB genotypesinimmunocompetent, immunocompromised, and congenitally infected Italian populations. Arch Virol. 2003;148(3):547-54. [ Links ]
10. Navarro D, Paz P, Tugizov S, Topp K, La Vail J, Pereira L. Glycoprotein B of human cytomegalovirus promotes virion penetration into cells, transmission of infection from cell to cell, and fusion of infected cells. Virology. 1993;197(1):143-58. [ Links ]
11. Cunha AA, Aquino VH, Mariguela V, Nogueira ML, Figueiredo LTM. Evaluation of glycoprotein B genotypes and load of CMV infecting blood leukocytes on prognosis of AIDS patients. Rev Inst Med Trop Sao Paulo. 2011;53(2):82-8. [ Links ]
12. Chou S, Dennison KM. Analysis of interstrain variation in cytomegalovirus glycoprotein B sequences encoding neutralization-related epitopes. J Infect Dis. 1991;163(6):1229-34. [ Links ]
13. Chakrabarti S, Mautner V, Osman H, Collingham KE, Fegan CD, Klapper PE, et al. Adenovirus infections following allogeneic stem cell transplantation:incidence and outcome in relation to graft manipulation, immuno suppression, and immune recovery. Blood. 2002;100(5):1619-27. [ Links ]
14. Ison MG. Adenovirus infections in transplant recipients. Clin Infect Dis. 2006;43(3):331-9. [ Links ]
15. Khanna R, Smith C. Cellular immune therapy for viral infections in transplant patients. Indian J Med Res. 2013;138(5):796-807. [ Links ]
16. Green M. Management of Epstein- Barr Virus-induced Post-transplant Lympho proliferative Disease in Recipients of Solid Organ Transplantation. Am J Transplant. 2001;1(2):103-108. [ Links ]
17. Mercorelli B, Lembo D, Palù G, Loregian A. Early inhibitors of human cytomegalovirus: state- of-art and therapeutic perspectives. Pharmacol Ther. 2011;131(3):309-29. [ Links ]
18. Drew WL,Chou S,Miner RC,Mohr BA,Busch MP,Van der Horst CM, et al. Cytomegalovirus glycoprotein B groups in human immunodeficiency virus- infected patients with incident retinitis. J Infect Dis. 2002;186(1):114-7. [ Links ]
19. Ahumada-Ruiz S, Taylor-Castillo L, Visoná K, Luftig RB, Herrero-Uribe L. Determination of human cytomegalovirus genetic diversity in different patient populations in Costa Rica. Rev Inst Med Trop Sao Paulo. 2004;131:87-92. [ Links ]
20. Lion T, Baumgartinger R, Watzinger F, Matthes- Martin S, Suda M, Preuner S, et al. Molecular monitoring of adenovirus in peripheral blood after allogeneic bone marrow transplantation permits early diagnosis of disseminated disease. Blood. 2003;102(3):1114-20. [ Links ]
5Abbreviations: BK virus, BKV; Caja Costarricense del Seguro Social, CCSS; Cytomegalovirus, HCMV; Epstein Barr Virus, EBV; Genotype 3, gB3; Herpes 6, HHV6; Hospital San Juan de Dios, HSJD; Kidney transplantation, KT; Hematopoietic progenitor cell or bone marrow transplant, BMT. Sources of support: laboratory tests for transplant patients, Molecular Biology Laboratory, Dr. Clodomiro Picado T. Clinical Laboratory.
Received: December 18, 2019; Accepted: August 19, 2021










 texto em
texto em