Introduction
Dental caries is one of the most prevalent diseases around the world, according to the World Health Organization about 2000 million people suffer from it (1), and over 530 million children suffer from caries in the primary dentition (2). In our country, the prevalence of dental caries in children has been hardly investigated; however, in 1999 an Oral health survey was conducted; it showed that the prevalence of caries in primary dentition was 75.2% in infants between 6 and 8 years old (3,4).
The progression of carious lesions is closely related to sugar consumption, oral hygiene habits, systemic conditions, and oral microbiota (5). According to Tanner et al. (6), this oral microbiota is an ecological community of commensal or symbiotic microorganisms and pathogens that share the same place in our body; it colonizes the dental surfaces naturally forming the biofilm.
The oral microbiota is composed of hundreds of bacterial species, where environmental factors (such as diet, type of calving, geographic location, and host genetics) play an important role in shaping the community structure of the microbiome and determining the pathogenic species associated with dental caries (7).
Traditionally, Streptococcus mutans was accredited as the main cause of dental caries; nonetheless, since the 2000s, there have been studies related to cultures strictly of carious lesions that have evidenced the presence of Gram-positive bacteria such as Lactobacillus, Streptococcus, Propionibacterium, Bifidobacterium, Actinomyces, Eubacterium, Rothia, Arachnia, Micromonas, Pseudoramibacterium; and Gram-negative bacteria like Prevotella, Porphyromona and Selenomones (8,9,10,11,12,13,14).
Recent research focused on pediatric population confirm this heterogeneity in the microbiota; e.g. in Columbus, Ohio, USA, Aas et al. (15) identified bacterial groups in carious lesions in infants and adolescents; some of them were Streptococcus mutans, Atopobium, Propionibacterium, Lactobacillus, Actinomyces spp. and Atopobium spp. In Canada, Agnello et al. (16) recognized a greater number of microbial communities like Veillonella, Porphyromonas, Streptococcus gordonii, and Strep- tococcus sanguinis. On the other hand, in European and African populations, Yang et al. (17) determined the presence of Actinobacteria and Firmicutes, Porphyromonas gingivalis, Prevotella intermedia, Treponema denticola, and Filifactor Alocis.
Due to the fact that the cariogenic microbiota constantly changes (based on the race or origin place of each person), and that in Central America, specifically Costa Rica, no studies have been performed regarding this field, the objective of this research was to identify the bacteria present in the microbiota of dentinal carious lesions in primary molars in Costa Rican pediatric patients.
Methodology
A convenience sample of 15 children aged between 4 and 8 years with deep carious lesions in primary molar dentin was used. The inclusion criteria were: infants who were actively attended by students at the Faculty of Dentistry (FD) from the UCR, and whose parents or legal guardians signed the informed consent to participate in this research. The exclusion criteria were: uncooperative patients for sample collection and children who are under chronic pharmacological treatments or during the last 3 months.
The sample collection was done as follow; the molar with advanced caries was selected in each patient; the operator applied an anesthetic technique based on that molar, and absolute isolation was placed with rubber dam sheet. Then, using a sterile spoon, the researcher removed a portion of the decayed dentin and deposited it directly in a sterile centrifuge tube with 2cc of trypticase soy broth (TSB). Finally, it was transpor- ted to the laboratory. After taking the sample, the operator continued with the restorative treatment of the decayed molar.
In less than 3 hours, this sample was transferred to the microbiology laboratory from the UCR, where it was placed in TSB for 24 hours in agitation, with a temperature of 37°C to guarantee a homogeneous liquid. After incubation, the broth was sown in Blood Agar; the plates were stored for 24 and 48 hours in the Carbon Dioxide (CO2) chamber (ThermoFisher Scientific, Waltham, Massachusetts, Estados Unidos) at 37°C to promote bacterial growth. Once this time had elapsed, the macroscopic characterization of the different bacterial colonies was carried out to determine their shape, size, and color. Each bacterial strain was isolated in a new plate of Blood Agar and stored for 24 hours in the CO2 chamber (ThermoFisher Scientific, Waltham, Massachusetts, Estados Unidos) at 37°C as in Elbing's and Brend's study (18).
After 24 hours, Gram stain were performed following the procedure implemented by Coico (19) to confirm the purity of each strain. At the end of the Gram stain drying, the sample was observed under the microscope (Olympus Life Science®, Waltham, Massachusetts, Estados Unidos) (20).
The bacterial strains identified as Gram- positive were examined with the catalase test as Taylor in 1972 mentioned (21). If the result was negative, the VITEK 2 (VITEK® bioMérieux, Ciudad de México, México) test described by Vargas et al. was performed (22). When the result of the catalase test was positive, a rapid effervescence with bubble detachment was formed; then, the bacterial identification test API STAPH (API® bioMérieux, Ciudad de México, México) (mentioned by Holmes et al. and bioMérieux) was applied (22,23,24).
In the case of the bacterial strains identified as Gram-negative, the oxidase test was performed, according to the method described by Bou (25), through strips of paper impregnated with the reagent para-amino-N-dimethyl-aniline (Bioser® S.A, Barcelona, España), and also the Triple Sugar Iron Agar test (T.S.I Agar), as González (26) applied it in his study to determine the fermentation capacity of bacteria with lactose, glucose or sucrose. When the oxidase test result was negative, bacterial identification was performed, specifically, the API 20E (API® bioMérieux, Ciudad de México, México) tests mentioned by Holmes et al. and bioMérieux (23,24). However, when the oxidase test was positive or there was no more than an 85% probability of identification, the VITEK 2 (VITEK® bioMérieux, Ciudad de México, México) bacterial identification test described by Vargas et al. was applied (22).
Results
Of the 15 samples of carious lesions in primary teeth, 60 pure bacterial strains were isolated through striatum in Agar plates. In each sample, a different number of strains was obtained, not only after 24 hours the growing was different but also after 48 hours because there are bacteria that take a longer time to cultivate.
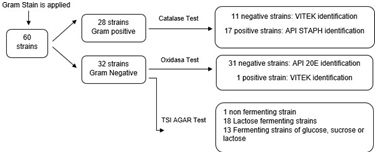
Of the 60 bacterial strains subjected to the Gram test, 28 Gram-positive and 32 Gram-negative strains were obtained. Of the Gram-positive strains, 11 were negative for the catalase test, so they were examined with VITEK 2 (VITEK® bioMérieux, Ciudad de México, México). In addition, the other 17 positive strains were analyzed with the API STAPH (API® bioMérieux, Ciudad de México, México), test for their bacterial identification. On the other hand, all 32 Gram-negative bacterial strains were tested with TSI, and the results were the following: only 1 non-fermenter (it did not change its color); 18 are only glucose fermenters; 13 fermented to glucose, sucrose or lactose. Also, the oxidase test was applied to these 32 stains, in which 31 were negatives, so the API 20E (API® bioMé- rieux, Ciudad de México, México), test was used, and the only positive strain of oxidase (Bioser® S.A, Barcelona, España), was identified through the VITEK 2 test (VITEK® bioMérieux, Ciudad de México, México) (Figure 1).
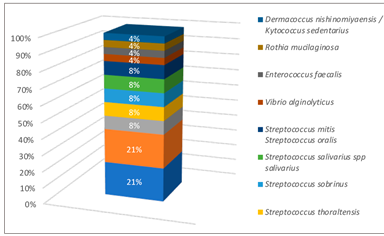
After obtaining the preliminary bacterial identification tests, the microorganisms present in the dental caries samples were determined through the biochemical methods: the quantity of bacterial strains identified with API STAPH test (API® bioMérieux, Ciudad de México, Mexico) was 12: 6 Staphylococcus epidermis, 2 Staphylococcus sciuri, 2 Staphylococcus xylosus and 2 Staphylococcus aereus. With API 20E test (API® bioMérieux, Ciudad de México, Mexico) it was obtained 22 strains: 6 PasteurellaPneumotropica/Mannheimia haemolytica, 1 Photobacterium damselae, 2 Serratia marcescen, 6 Pantoea spp, 1 Klebsiella pneumoniae spp azaenae, 1 Pseudomonas fluorescens putida and 1 Shigella spp. Finally with VITEK 2 (VITEK® bioMérieux, Ciudad de México, Mexico) a total of 24 strains: 5 Streptococcus mutans, 2 Streptococcus sanguinis, 2 Streptococcus thoraltensis, 2 Streptococcus sobrinus, 2 Streptococcus salivarius spp salivarius, 2 Streptococcus mitis Streptococcus oralis, 1 Vibrio alginolyticus, 1 Enterococcus fecalis, 5 Sphingomonas paucimobillis, 1 Rothia mucilaginosa and 1 Dermacoccus nishinomiyaensis/Kytococcus seden- tarius. The results in percentages can be seen in Figure 2, Figure 3 and Figure 4.
Discussion
The diversity of the bacterial microbiota is a very relevant issue (16,27,28). This research strengthens the evidence not only the Streptococcus mutans are isolated in advanced caries lesions, but also there are different bacterial strains that act together to destroy dental tissues.
Gram-positive bacteria were found in this research, whereby Streptococcus mutans previously identified as specific bacteria of dentinal caries in primary pieces (29) was present in some samples. This result agreed with the study made by Quidemat et al. (30), in which they reported that some subjects with active caries had detectable levels of Streptococcus mutans bacteria. However, the researchers also pointed out a high level of anaerobic bacteria such as Leptotrichia shahii and Prevotella melaninogenica, but this situation did not occur in the present study because only aerobic bacteria were identified. Other Streptococcus found were: Streptococcus sanguinis and Streptococcus salivarius, situation also presented in Bansal's research (31).
Regarding the Staphylococcus, the main subvariant presented was the Staphylococcus epidermidis, similar to the research of Devang et al. (32); they indicated that this microorganism has a high adaptive capacity, which allows it to obtain, lose or regulate genetic elements providing better environments for colonization in the host. That is why, it is so prevalent in dental caries.
An important finding in this research was the presence of Enterococcus faecalis which is a Gram-positive; facultative anaerobic coccus bacterium that possesses numerous virulence factors, according to Carrero et al. (33). It may be found transiently in oral cavity and soft tissue lesions, principally in populations exposed to hospital environments or in nosocomial infections; For this reason, it could be assumed that children might be exposed to these contexts prior to the sampling appointment.
Another Gram-positive bacterium found in a small number of strains in this study was Rothia mucilaginosa; is part of respiratory tract and oral cavity, but also has the ability to produce several infections and dental caries progression combined with other microorganisms (34,35). Rothia mucila- ginosa that was not observed in the research of De Jesus et al. (36), who instead reported the presence of bacteria like Mycosphaerella, Cyber- lindnera, and Trichosporon, in samples of deep carious lesions. Other studies also reinforce the heterogeneity of the oral microbiota; even though they did not report the finding of Rothia, they identi- fied Veillonella and Actinomyces. These studies were carried out in the United States and Germany, in several age populations, so they did not share either geographical location or age range (37,38). Rothia mucilaginosa has been classified an opportunistic pathogen, it has the ability to be found in symbiosis and dysbiosis, which is why it was able to detect in deep caries in this investigation (34).
Regarding Gram-negative bacteria, Pantoea was identified. This is an enterobacterium frequently isolated in plants, fruits, and vegetables, accor- ding to Segado et al. (39), so the presence of this bacterium could be due to the fact that some of the subjects (before receiving dental care) did not perform proper oral hygiene; therefore, it is possible that food residues are lodged in an open cavity of caries.
They also identified the strains Sphingomo- nas paucimobilli and Pasteurella pnuemotropica, both Gram-negative. It is a relevant finding to highlight in this research because Sphingomonas paucimobilli is a bacterium located in aqueous media and is rare to find in humans; in addition, its persistence is related to infections and fever in pediatric patients (40). On the other hand, Pasteurella pnuemotropic is an opportunistic pathogen presented in the oral cavity of rodents, dogs, and cats. In fact, there are cases of mortality due to septicemia and long stays in hospitals that have been reported (41,42,43). In other words, the results in this research could be considered an isolated event, and not directly related to carious lesions in dentin; however, it can be assumed that children have been exposed to environments with any of the aforementioned animals; therefore, the presence of this bacterium.
The bacterial heterogeneity identified in this research accorded with studies which have established that the microbiota varies based on its location in different dental structures. It also indicates that the destruction in tissues from enamel to dentin involves different microorganisms in each area. Streptococcus, Veillonella and Actinomyces have been identified in carious enamel lesions. While the destruction increases and caries is harbored in dentin, other bacteria such as Prevotela denticolens and Lactobacillus can be detected (29,37,44) .
One of the benefits of this study is that it was the first research which identified the microbiota of carious lesions in dentin of primary teeth of Costa Rican children establishing similarities and differences with others, in several countries (29,31,38). However, one of the limitations was that the same identification methods were not used. In Umea Sweden, the detection was performed by Polymerase Chain Reaction (PCR) in samples obtained in 13 children, and the main species associated with caries were Actinobaculum, Atopobium, Aggregatibacter and Streptococcus mutans (38). In addition, Ribeiro et al. (45) applied 16S and n rRNA metage- nomic identification in 13 children from Rio de Janeiro, Brazil; they found Streptococcus, Coryne- bacterium matruchotii, Actinomyces viscosus Prevotella nigrescens and Dialister micraerophilus.
One of the forward-looking statements of this research is to continue with the characterization of the microbiota using molecular identification techniques, and to relate the presence of different types of bacteria with other variables such as diet type and oral hygiene.
Conclusion
The bacteria of the oral microbiota of Costa Rican pediatric patients, with advanced stages of dentinal carious lesions in primary dentition, which were presented in a greater number of strains, were Staphylococcus epidermidis, Pasteurella ppneumotropic/Mannheimia haemolytica, Pantoea spp, and Streptococcus mutans.
Author contribution statment
Conceptualization and design: K.T.S., N.G.M., T.R.M. and P.A.S.
Literature review: K.T.S.
Methodology and validation: K.T.S., N.G.M., T.R.M. and P.A.S.
Formal analysis: N.G.M., T.R.M. and P.A.S. Research and data collection: K.T.S.
Resources: K.T.S. and P.A.S.
Data analysis and interpretation: K.T.S., N.G.M.,T.R.M. and P.A.S.
Writing and preparation of the original draft: K.T.S. and N.G.M.
Writing, proofreading and editing: K.T.S., N.G.M., T.R.M. and P.A.S.
Supervision: N.G.M., T.R.M. and P.A.S. Project management: .T.S. and N.G.M.
Acquisition of funds: Not applicable for this study.