Introduction
The family Calliphoridae of blow flies is among the families most studied in the world. This family has synanthropic habitats that impact directly in forensic entomology, medical, veterinary and economic subjects (3, 4, 5 y 6). In particular, blow flies are among the first colonizers of cadavers that might serve as a biological clock to measure the time of death and are important in an estimation of the post-mortem interval (PMI) (3). More than 1000 species are described for Calliphoridae; 126 species are found in the Neotropics, belonging to subfamilies Calliphorinae, Chrysomyinae, Melanomyiniae and Luciliinae (7). Specifically, as the subfamily Luciliinae is diverse and heterogeneous, taxonomic classification becomes confusing (8). As many taxonomic descriptions for Luciliinae blow flies lack sufficient resolution, it is difficult to verify the species that they represent without seeing the original types (6). Inside Luciliinae, the genus Lucilia is forensically important; shortcomings have been described also for taxonomical identification leading to confusion and difficulty in resolving the species status for any but the most common species (6). Accuracy is mandatory in the identification of forensically important Lucilia species, but can be difficult because of similarities in their morphology (9); only experts such as taxonomists and trained technicians can identify taxa accurately through extensive experience (10). As a DNA approach was proposed for the identification of insect species for forensic issues (11) and for species identification (12), cytochrome oxidase I (COI) has been used world-wide for molecular identification of forensically important species (9, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27)such as Calliphoridae and Sarcophagidae, provides evidence for estimation of postmortem interval (PMI, including blow-fly genus Lucilia (28, 29, 30, 31). Some Lucilia species are so cosmopolitan that forensic entomology is specific to a locality; molecular studies are generally conducted in particular regions. The DNA of specimens sequenced from new localities might not exactly match published DNA sequences, indicating levels of variation, cryptic complex or revealing paraphyletic patterns of studied species (10, 19). In particular, in Costa Rica, Whitworth (6) described morphologically and molecularly (using COI) specimens of Lucilia cuprina (1), L. eximia (1), L. cluvia (2), L. rognesi (6), L. woodi (6), L. pulverulenta (6) and L. purpurascens (2). This extensive work, was conducted with samples collected at several locations in Costa Rica, provided not only new taxonomic keys for Lucilia but also vouchered sequences that we used to compare our DNA sequences. Our work seeks to analyze 42 Lucilia specimens in a new set collected in Costa Rica, of which 21% correspond to flies reared from larvae taken from human cadavers. The identification of these individuals provides us with valuable information about the principal species involved in the decomposition processes that are relevant for forensic purposes.
Materials and methods
Sampling and taxonomic identification of Lucilia species.
Adult specimens (n = 42) were obtained from a dry reference collection of the Department of Forensic Science, Costa Rica. Information on the collection sites and sampling method (where available) was annotated (data not shown). Because of the antiquity of the collection, information about the sampling method is, unfortunately, not available for all samples. For cases in which larvae were taken from human corpses, they were reared to adulthood; these adults were used in our morphological and genetic analysis. Specimens were morphologically classified by a forensic entomologist according to taxonomic keys of Vargas (32), Morales (33) and Whitworth (6).
DNA extraction and PCR amplification
Two legs of each blow fly were used for DNA isolation (DNeasy Blood & Tissue kit, QIAGEN, final elution volume 70 µL) according to the manufacturer´s instructions. Two regions were partially amplified with PCR: (i) cytochrome c oxidase subunit I (COI) and (ii) 28S ribosomal subunit from nuclear ribosomal DNA. PCR reactions for the COI region were conducted in a final volume 20 µL, containing HotStart Ready Mix (1X, Fermentas), primer LCO1490-L (5′-GTCWACWAATCATAAAGATATTGG-3′; 0.4 mM), primer HCO2198-L (5′-TAAACTTCWGGRTGWCCAAARAATCA-3′; 0.4 mM) and the extracted DNA (2.5 µL in the range 3-15 ng/µL). PCR was performed with an initial denaturation step of 95 °C for 5 min, followed by 35 cycles of 94 °C for 30 s, 45 °C for 30 s and 72 °C for 2 min, and a final elongation step of 72 °C for 5 min. PCR reactions for the 28S region were conducted with the same volumes of COI reactions and with primers 28S-F1 (5´-GGGAGGAAAAGAAACTAACAAGG-3´) and 28SR1 (5´-CTGTTTCGGTCTTC CATCAGGG-3´) (4). PCR reactions were performed with an initial denaturalization step of 95 °C for 5 min, followed by 35 cycles of 94 °C for 45 s, 52 °C for 60 s and 72 °C for 2 min, and a final elongation step of 72 °C for 5 min. A 96-well thermal cycler (Veriti, Applied Biosytems) was used for both PCR reactions. PCR products (5 µL) were purified (Exonuclease I, 1 µL, Thermo Scientific Fermentas) and Phosphatase Alkaline (0,5 µL, Thermo Scientific Fermentas). The mixtures were placed in a 96-well thermal cycler (Veriti, Applied Biosytems) with initial temperature 37 °C for 15 min, followed by 85 °C for 15 min. Sequencing reactions were conducted with the BigDye Terminator Kit (Applied Biosystems) with final primer concentrations 0.320 µM of either forward or reverse primer. The sequencing reactions were performed in a 96-well thermal cycler (Veriti, Applied Biosytems) with the following program: 1 cycle of initial temperature 96 °C for 1 min; 15 cycles of 96 °C for 10 s, 50 °C for 5 s and 60 °C for 75 s; 5 cycles of 96 °C for 10 s , 50 °C for 5 s and 60 °C for 90 s; 5 cycles of 96 °C for 10 s, 50 °C for 05 s and 60 °C for 2 min. Sequencing products were purified with a kit (BigDye XTerminator®, Applied Biosystems) according to the manufacturers instructions. Purified sequencing reactions were analyzed in a genetic analyzer (3130xl, Applied Biosystems).
Sequence and phylogenetic analyses
Forward and reverse strands were manually inspected (FinchTV, Geospiza); a consensus sequence was obtained in BioEdit (34). Consensus sequences were compared on the NCBI through the function Nucleotide BLAST (Basic Local Alignment Search Tool) to identify the homology of the sequences obtained. Multiple sequence alignments of the COI and 28S regions were conducted in Mega version 6 (35) with MUSCLE option (36) using sequences from this work and also from databases GeneBank and BOLDSYSTEMS. Models of nucleotide substitution were tested in MEGA6 (35); Tamura 3 parameters were selected to build the neighbor-joining (NJ) tree with 1000 bootstrap replicates.
Results and Discussion
Morphological identification of Lucilia species In total, three species of forensic interest belonging to the subfamilie Luciliinae were identified morphologically: L. cuprina (33.3 % of total specimens), L. eximia (33.3 %) and L. purpurascens (21.4 %) (figure 1 and table 1). These species are known in Costa Rica and are distributed throughout the entire country (6, 33). We identified also one cryptic complex assigned as Lucilia Complex (12.0 % of total specimens). Specimens included in this complex have phenotypical characteristics that allowed the distinction from specimens of L. cuprina, L. eximia and L. purpurascens, but we could not classify samples from the Complex all the way to the species. A new taxonomical key is reported in which new species, including L. rognesi, L. woodi, L. pulverulenta and L. cluvia, are described for the Neotropic region (6). Despite these findings, some characteristics in the new taxomic key were not clearly identified in our samples; we hence kept specimens as Lucilia Complex.
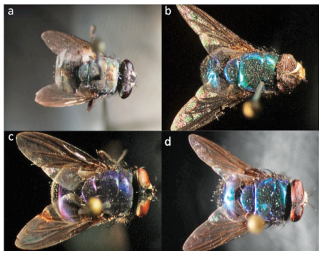

Figure 1 Taxonomic classification of specimens included in this study a) Lucilia cuprina, b) Lucilia eximia, c) Lucilia purpurascens and d) Lucilia Complex
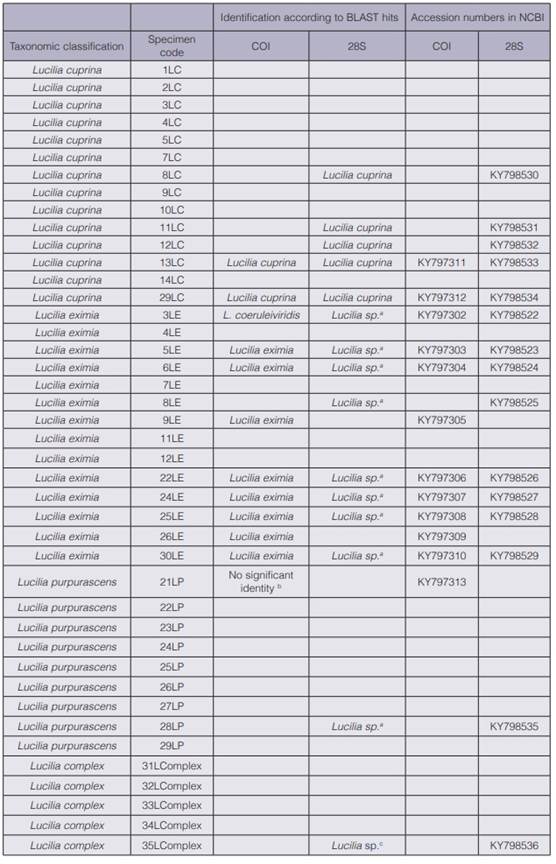
Table 1 Taxonomic classification of 42 Lucilia specimens collected in Costa Rica and partial molecular identification with COI and 28S regions.
Alignment of COI and 28S sequences
DNA extraction was possible for all samples (42 specimens), but not all DNA were amplified with either COI or 28S. In total 12 samples were amplified for COI and 15 for 28S; all sequences were deposited in the GeneBank (table 1). This result represents rates 28.6 % and 35.7 % of PCR amplification success, respectively. Of our specimens most were collected several years ago (data not shown) and pinned for taxonomic studies; this factor degrades the DNA. Moreover, phenolic compounds were used for preservation. One limitation in the DNA barcoding surveys is the condition of a sample (10). For this reason this method has been used mainly on only fresh specimens or samples preserved in an ideal manner for molecular work (i.e. refrigerated, stored in ethanol or acetone and sampled in liquid nitrogen) (10, 16, 26, 30, 37). BLAST results showed that, of nine samples morphologically classified as L. eximia amplified for COI, eight had significant identity to L. eximia with no intra-specific sequence variability. Sample 3LE was taxonomically classified as L. eximia but the sequence alignment had similarity to Lucilia coeruleiviridis (table 1). For specimens morphologically classified as L. cuprina, two amplified for COI with no variability in the sequences; alignment results in BLAST identified this species accordingly. Only one sequence was obtained for L. purpurascens that lacked significant identity with any Lucilia species (table 1). The 28S region lacked sufficient resolution to identify samples to the level of species except for L. cuprina (table 1). Sequences of L. eximia, L. purpurascens and Lucilia Complex showed identities to only the Lucilia genus level. The sequences of 28S for L. eximia, L. purpurascens and Lucilia Complex aligned with species such as L. mexicana (38) , L. sericata (39) and L. cluvia. The only sequence of Lucilia Complex (35L Complex) for 28S aligned for other species such as L. thatuna (40), L. coeruleiviridis (38), L. infernalis and L. porphyrina (2) (table 1). The difference in resolution when using COI and 28S was reported by Zajac et al. (27), who found that identification of forensically relevant Diptera species in Thailand appeared to be more conclusive with COI than with 28S. This region failed to differentiate the two closely related species Lucilia illustris and Lucilia caesar (29). Despite 28S being inconclusive, our work is a contribution to sequence databases for future Lucilia DNA barcoding studies world-wide. To date only few sequences for solely 28S are available for L. eximia (4).
Phylogenetic analyses with COI and 28S regions
The inferred phylogenetic tree obtained with NJ (figure 2) shows the analysis for COI in which all samples of L. cuprina grouped with the reference sequences. The bootstrap values confirmed that molecular identification of L. cuprina is not complex and can be easily assayed with COI (figure 2), but identification of L. cuprina with conventional taxonomy was also congruent with molecular taxonomy. No monophyletic pattern using COI has been reported for L. cuprina (19, 25, 28, 41). DeBry et al. (30) found a monophyletic group for L. cuprina in a vouchered collection of Lucilia species identified morphologically. We obtained two sequences for L. cuprina (13LC and 29LC) without intra-specific variation with reference sequence ASIN3389-12 (figure 2) collected in Costa Rica. The departures from monophyly can be detected only with replicate samples; additional sequences are required before the COI monophyly can be known. L. cuprina has shown morphological variation according to its geographic distribution (14). The NJ tree for COI region grouped almost all our L. eximia samples (figure 2) with specimens collected in Costa Rica (ASIND3390-12, ASIND3391-12) and those described by Whitworth (6); the exception was sample 3LE that grouped with accessions belonging to species newly described by Withworth (6). This group contains also a reference accession classified as L. eximia from Costa Rica (ASIND3394-12). Specimens of L. eximia have been found to group into several discrete clusters widely separated according to geographical region; the results hence raise the possibility that L. eximia is a cryptic species (6). As mentioned above, the BLAST result of sample 3LE indicated an alignment with L. coeruleiviridis that is impossible because this species has not been found in the Neotropical region (6). A NJ analysis was run with reference sequences of L. coeruleiviridis from publications of Wells et al. (28), DeBry et al. (30) and Whitworth (6). Sample 3LE kept grouped according to figure 2, but reference samples of L. coeruleiviridis also located in the same group. This result indicates a revision of conventional taxonomy considering that specimens classified as L. eximia might belong to a new species (L. rognesi, L. woodi, L. pulverulenta) in Costa Rica, or as stated above reflects the possibility of a cryptic species. The only sample classified as L. purpurascens (21LP) grouped with no reference accession (figure 2), despite our inclusion as references of several accessions of L. purpurascens with synonymy L. purpurEscens, L. peruviana (6).
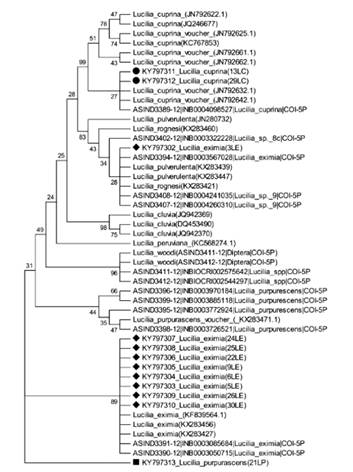
Figure 2 Phylogenetic tree of Lucilia specimens in this study and sequences from databases. The Neighbor-Joining method (with 1000 bootstrap replicates) was used with the COI region. The evolutionary distances were computed with the Tamura 3-parameter method and have as unit the number of base substitutions per site.
Despite samples 3LE and 21LP displaying differentially in the NJ tree, we do not declare our specimens as distinct species because more specimens of male and female blow flies are required for classification with conventional taxonomy, but the use of COI increases the accuracy for the identification of Lucilia species; additional knowledge other than genotype is required to shorten the list of candidate species to include only forms that can be distinguished with COI (18, 28). The absence of concordance between morphological and molecular taxonomy in samples 3LE and 21LP revealed the great complexity and the unresolved problem with Lucilia classification with conventional taxonomical keys, as occurs analogously with another Lucilia species (20, 30, 31). The 28S region showed no resolution to identify some samples when sequences were aligned in GeneBank databases, except L. cuprina (table 1). As expected, the NJ tree (figure 3) revealed a monophyletic group with L. cuprina, as similarly described by Debry et al. (41). A second group contained L. eximia (including sample 3LE) and our specimens classified as L. purpurascens (21LP) and Lucilia Complex (35LComplex).

Figure 3 Phylogenetic tree of Lucilia specimens in this study and sequences from databases. The Neighbor-Joining method (with 1000 bootstrap replicates) was used with the 28S region. The evolutionary distances were computed using the Tamura 3-parameter method and have as unit the number of base substitutions per site.
L. eximia and L. cuprina are relevant species for possible forensic applications in Costa Rica The joint analysis of the morphological and molecular data allows us to conclude that 66 % of the specimens of the OIJ collection belong to two species - L. eximia and L. cuprina. To our knowledge, the morphological and molecular analyses have not been reported for L. eximia and L. cuprina found in human cadavers, which points to these two species being two of the most relevant for forensic purposes in Costa Rica. For these two species, the necessary requirements are met for subsequent forensic application, as (i) it is possible to find larvae in human bodies, (ii) larvae can be grown to adulthood, and (iii) both morphological and molecular analyzes allow the identification of these species appropriately. Large sets of Calliphoridae samples have been studied around the world with specimens collected in varied environments and with various collection methods (i.e. nets, meat bait, liver-baited traps, pig carcases) that have increased the knowledge of the taxonomy in this family (6, 9, 13, 14, 19, 20, 30). Larvae samples collected from human cadavers have also been reported but in a minor proportion. In Portugal, the blowfly species L. sericata, L. caesar Linnaeus, L. ampullacea Villeneveu and L. illustris (39) were identified when maggots (or reared larvae) were obtained from autopsies on human cadavers (31, 42, 43). Species L. sericata was found in samples from corpses in Germany (22).
Conclusion
Despite Whitworth (6) described Lucilia species from Costa Rica, our work enriches the knowledge of forensically important Lucilia species in Costa Rica. Most samples were accordingly classified with morphology and molecular analysis with COI or 28S regions, but we found contradictions for two samples (3LE and 21LP). The contradictions in morphological and molecular analyses points to the necessity of developmental characters related with the varied microclimates and geography of Costa Rica. The results of morphological and molecular analysis revealed that larvae collected from human corpses and reared to adulthood correspond to only L. cuprina and L. eximia. We identified also individuals from L. purpurascens and Lucilia complex. To our knowledge, this work is the first in Costa Rica and the Centro-American region to describe cadaveric entomofauna, which is valuable for the development of potential forensic applications in estimating the postmortem intervals. This work provides additional DNA sequences for Lucilia flies that increases the knowledge of the genetic diversity in this genus.
Acknowledgments
Centro Nacional de Innovaciones Biotecnológicas (CENIBiot) and Tecnológico de Costa Rica (TEC) supported the experimental work of this study. S.S-A received funding for a Student Research Project by TEC. We thank Manuel Zumbado, John Vargas and Terry Whitworth for providing information about the taxonomy of Lucilia species that helped us to address our work.